https://reasonandscience.catsboard.com/t2234-the-origin-of-replication-and-translation-and-the-rna-world
A self-replicating molecule has never been seen. But also if it existed, it would be helpless to create a living cell. If molecule A self-reproduces n-times we would have AAAAAAA....That is ridiculously trivial and has nothing to do with what we see in a cell. A cell is a cybernetic ultra-complex system, where, thanks to countless concurrent software-driven chemical and physical processes using languages and codes, material is stored, managed, moved, assembled, converted and positioned such that the cell survives and self-reproduces. To believe, as abiogenesis and evolutionists do, that AAAAAAA... leads to a cell, is like to think that by simply duplicating bricks BBBBBBB... we get a functioning complete nuclear plants.
Martina Preiner: The Future of Origin of Life Research: Bridging Decades-Old Divisions 2020 Feb 26
Many found the metaphor appealing: a world with a jack-of-all-trades RNA molecule, catalyzing the formation of indispensable cellular scaffolds, from which somehow then cells emerged. Others were quick to notice several difficulties with that scenario. These included the lack of templates enabling the polymerization of RNA in the prebiotic complex mixture and RNA’s extreme lability at moderate to high temperatures and susceptibility to base-catalyzed hydrolysis
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7151616/?fbclid=IwAR0RQGnrvfMnW3qaQeRHIYtAdVwyS3KhBjoLlAjAOccF0H9-Phgv7acaME0
The phrase "RNA World" was first used by Nobel laureate Walter Gilbert in 1986, in a commentary on how recent observations of the catalytic properties of various forms of RNA fit with this hypothesis.
https://en.wikipedia.org/wiki/RNA_world
Leslie Orgel: Prebiotic chemistry and the origin of the RNA world Mar-Apr 2004
It is possible that all of these, and many other difficulties, will one day be overcome and that a convincing prebiotic synthesis of RNA will become available. However, many researchers in the field, myself included, think that this is unlikely
https://pubmed.ncbi.nlm.nih.gov/15217990/
Christian de Duve: The Beginnings of Life on Earth September-October 1995 issue of American Scientist.
Contrary to what is sometimes intimated, the idea of a few RNA molecules coming together by some chance combination of circumstances and henceforth being reproduced and amplified by replication simply is not tenable. There could be no replication without a robust chemical underpinning continuing to provide the necessary materials and energy.
https://www2.nau.edu/~gaud/bio372/class/readings/beglifeerth.htm
Robert Shapiro: Small Molecule Interactions were Central to the Origin of Life (June 2006),
The myth of a small RNA molecule that arises de novo and can replicate efficiently and with high fidelity under plausible prebiotic conditions . . . [is] unrealistic in light of current understanding of prebiotic chemistry
https://www.jstor.org/stable/10.1086/506024
W.Gilbert: The RNA world 20 FEBRUARY 1986
If there are activities among RNA enzymes or ribozymes, that can catalyze the synthesis of a new RNA molecule from precursors and an RNA template, then there is no need for protein enzymes at the beginning of evolution. One can contemplate an RNA world, containing only RNA molecules that serve to catalyze the synthesis of themselves. The first stage of evolution proceeds, then, by RNA molecules performing the catalytic activities necessary to assemble themselves from a nucleotide soup. The RNA molecules evolve in self-replicating patterns, using recombination and mutation to explore new functions and to adapt to new niches. At the next stage, RNA molecules began to synthesize proteins, first by developing RNA adapter molecules that can bind activated amino acids and then by arranging them according to an RNA template using other RNA molecules such as the RNA core of the ribosome. This process would make the first proteins, which would simply be better enzymes than their RNA counterparts. Finally, DNA appeared on the scene, the ultimate holder of information copied from the genetic RNA molecules by reverse transcription. After double-stranded DNA evolved there exists a stable linear information store, error-correcting because of its double-stranded structure but still capable of mutation and recombination. RNA is then relegated to the intermediate role that it has today - no longer the center of the stage, displaced by DNA and the more effective protein enzymes.
https://www.nature.com/articles/319618a0.pdf
My comment: So far, the storytelling goes. But is it true?
Neeraja Sankaran: Revisiting the RNA World with its inventor September 6, 2017
The RNA World Hypothesis is a model for the early evolution of life on earth proposed in 1986 by the molecular biologist Walter Gilbert, in which he posited that the earliest forms of life were likely composed entirely of RNA molecules. According to this scenario, the two fundamental functions of life, namely metabolism, and the closely associated process of catalysis, and replication, namely the passing on of the information about various activities to the next generation of offspring, performed in contemporary living systems by proteins and DNA respectively, were both carried out by RNA.
Today, thirty years after the RNA World was first proposed, no one has seen an actual living system that is completely based in RNA. Nevertheless, the hypothesis lives on in the origins of life research community, albeit in a hotly debated, highly contentious atmosphere. Although there are strong opponents, many researchers agree that although far from complete, it remains one of the best theories we have to understand “the backstory to contemporary biology.” Gilbert himself expressed some disappointment that “a self-replicating RNA has not yet been synthesized or discovered” in the years since he predicted his hypothesis, but he remains optimistic that it will emerge eventually.
https://atlasofscience.org/revisiting-the-rna-world-with-its-inventor/#:~:text=The%20RNA%20World%20Hypothesis%20is,composed%20entirely%20of%20RNA%20molecules.
My comment: So, nobody has been able to confirm the hypothesis in 30 years. And there are reasons for that.
PHILIP BALL: Flaws in the RNA world 12 FEBRUARY 2020
Self-replicating RNA may lack the fidelity needed to originate life
The hypothesis of an ‘RNA world’ as the font of all life on Earth has been with us now for more than 30 years, the term having been coined by the biologist Wally Gilbert in 1986. You could be forgiven for thinking that it pretty much solves the conundrum of how the replication of DNA could have avoided a chicken-and-egg impasse: DNA replication requires protein enzymes, but proteins must be encoded in DNA. The intermediary RNA breaks that cycle of dependence because it can both encode genetic information and act catalytically like enzymes. Catalytic RNAs, known as ribozymes, play several roles in cells.
It’s an alluring picture – catalytic RNAs appear by chance on the early Earth as molecular replicators that gradually evolve into complex molecules capable of encoding proteins, metabolic systems and ultimately DNA. But it’s almost certainly wrong. For even an RNA-based replication process needs energy: it can’t shelve metabolism until later. And although relatively simple self-copying ribozymes have been made, they typically work only if provided with just the right oligonucleotide components to work on. What’s more, sustained cycles of replication and proliferation require special conditions to ensure that RNA templates can be separated from copies made on them. Perhaps the biggest problem is that self-replicating ribozymes are highly complex molecules that seem very unlikely to have randomly polymerised in a prebiotic soup. And the argument that they might have been delivered by molecular evolution merely puts the cart before the horse. The problem is all the harder once you acknowledge what a complex mess of chemicals any plausible prebiotic soup would have been. It’s nigh impossible to see how anything lifelike could come from it without mechanisms for both concentrating and segregating prebiotic molecules – to give RNA-making ribozymes any hope of copying themselves rather than just churning out junk, for example.
In short, once you look at it closely, the RNA world raises as many questions as it answers. The best RNA polymerase the researchers obtained this way had a roughly 8% chance of inserting any nucleotide wrongly, and any such error increased the chance that the full chain encoded by the molecule would not be replicated. What’s more, making the original class I ligase was even more error-prone and inefficient – there was a 17% chance of an error on each nucleotide addition, plus a small chance of a spurious extra nucleotide being added at each position.
These errors would be critical to the prospects of molecular evolution since there is a threshold error rate above which a replicating molecule loses any Darwinian advantage over the rest of the population – in other words, evolution depends on good enough replication. Fidelity of copying could thus be a problem, hitherto insufficiently recognized, for the appearance of a self-sustaining, evolving RNA-based system: that is, for an RNA world.
Maybe this obstacle could have been overcome in time. But my hunch is that any prebiotic molecule will have been too inefficient, inaccurate, dilute and noise-ridden to have cleared the hurdle.
https://www.chemistryworld.com/opinion/flaws-in-the-rna-world/4011172.article
The difficult case of an RNA-only origin of life AUGUST 28 2019
Despite a large body of evidence supporting the idea that RNA is capable of kick-starting autocatalytic self-replication and thus initiating the emergence of life, seemingly insurmountable weaknesses in the theory have also been highlighted. Despite advances in prebiotic chemistry, it has not yet been possible to demonstrate robust and continuous RNA self-replication from a realistic feedstock. RNA in isolation may not be sufficient to catalyse its own replication and may require help from either other molecules or the environment.
https://portlandpress.com/emergtoplifesci/article/3/5/469/220563/The-difficult-case-of-an-RNA-only-origin-of-life
Eörs Szathmáry: Toward major evolutionary transitions theory 2.0 April 2, 2015
Despite recent progress, we still have no general RNA-based replicase that could replicate a great variety of sequences, including copies of its own.
https://www.pnas.org/content/112/33/10104
Tan; Stadler: The Stairway To Life:
RNA replication experiments require a supply of pure homochiral nucleotides, and a supply of Qβ replicase, an enzyme that is actually responsible for combining the nucleotides into RNA molecules. An abiotic origin for Qβ replicase is absurd because it consists of a combination of four protein subunits and more than 1,200 amino acids in a specific sequence—an enzyme of great complexity.
B.Alberts: Molecular Biology of the Cell
Although self-replicating systems of RNA molecules have not been found in nature, scientists are hopeful that they can be constructed in the laboratory.
https://www.ncbi.nlm.nih.gov/books/NBK26876/
Stephen C.Meyer, The return of the God hypothesis, page 218
The RNA-world hypothesis presupposes but does not explain, the origin of sequence specificity or information in the original functional RNA replicators. To date, scientists have been able to design RNA catalysts that will copy only about 10 percent of themselves.31 For strands of RNA to perform even this limited self-replication function, they must have very specific arrangements of their constituent nucleotide building blocks. Further, the strands must be long enough to fold into complex three-dimensional shapes (so-called tertiary structures). Thus, any RNA molecule capable of even limited function must have possessed considerable specified information content. Yet explaining how the building blocks of RNA arranged themselves into functionally specified sequences has proven no easier than explaining how the constituent parts of DNA might have done so. As de Duve noted in a critique of the RNA-world hypothesis, “Hitching the components together in the right manner raises additional problems of such magnitude that no one has yet attempted to do so in a prebiotic context.”
In 1971, Nobel Laureate Manfred Eigen described how the length of a prebiotic information molecule like RNA must fundamentally be limited according to the error rate during replication. Reproduction of longer molecules introduces more errors, and too many errors would lead to an exponential increase in errors over many generations—an error catastrophe. Eigen observed that living organisms require error correction during replication to avoid error catastrophe in long DNA molecules. Yet the error-correction mechanisms themselves must be coded in the same lengthy DNA molecules. This led to Eigen’s paradox: a self-replicating molecule faces a practical size limit of about one hundred nucleotides unless there are error-correction systems, but the error-correction systems themselves must be coded in molecules that are substantially longer than the practical limit. Eigen’s paradox only addresses errors during replication and the need for error-correction mechanisms during replication. The additional burden of accumulated molecular damage from radiation, oxidation, alkylation, chemical mutagens, pathogens, and water, especially over deep time, and the associated requirement for additional molecular-repair mechanisms, greatly compounds Eigen’s paradox.
What we need for abiogenesis is a self-replicating RNA that has just the right error rate during reproduction, resulting in information gain over millions of years with a replication rate that somehow improves over time and the preservation of information via repair mechanisms that spontaneously appear. What we have is the absence of a self-replicating molecule, evidence that replication processes prefer shortening and simplification of RNA over time, known instability of RNA over time, and no hope of arriving at the required repair mechanisms. All empiric observations tell us that organic molecules, once life has ended, rapidly devolve into a useless mixture, referred to as “asphalt.” How, then, can we rationally expect organic molecules to progress naturally toward complexity and organization in the absence of life?
The most successful ribozyme for RNA reproduction to date, reported by Attwater and colleagues in 2018, is actually a combination of two RNA molecules—one composed of 135 ribonucleotides and the other composed of 153 ribonucleotides. The most successful ribozyme is therefore quite complex, requiring the duplication of two RNAs, not just one. The appearance of the first copy of these complex molecules must also be explained by natural abiotic processes. Such processes strictly cannot include any form of chemical evolution because that is the very process we are trying to initiate. Let’s assume that Attwater’s ribozyme (the most successful ribozyme to date) was able to copy itself exponentially, so all that is needed is a natural explanation for the arrival of the two RNA molecules in the ribozyme. Let’s make it even simpler by requiring only the arrival of the smaller RNA molecule, containing 135 ribonucleotides. Let’s also assume that we have an endless supply of activated, concentrated ribonucleotides; that the ribonucleotides are homochiral and they spontaneously link together via the desired bonds to produce RNA; and after the growing chain reaches 135 ribonucleotides, the molecule stops growing. What would it take under these very generous assumptions to find the 135 ribonucleotide RNA needed for Attwater’s ribozyme? The chance of arriving at the correct RNA is one in 4^135, or one chance in 1.9 X 10^81. This is similar to the total number of atoms in the known universe. Creation of one copy of each of 1.9 X 10^81 possible RNA molecules, each containing 135 ribonucleotides, would therefore require more material than the mass of the universe. Hopefully, this places a rational perspective on the chances of obtaining a self-replicating RNA. You might be wondering: If two copies of a ribozyme are required to initiate a duplication process (one to act as the template and the other to act as the ribozyme), why wouldn’t the process that provided two copies simply produce lots of copies, so we don’t need this type of reproduction process after all? That is a good question. The answer, of course, is that you can’t get the original two copies through natural abiotic processes in the first place—especially if each copy is a complex molecule such as a combination of 135 ribonucleotides and 153 ribonucleotides.
A highly touted report of an RNA ribozyme that could reproduce itself appeared in 2009. The title “Self-Sustained Replication of an RNA Enzyme” seemed to proclaim proof for abiotic reproduction of RNA. However, three serious limitations cast a shadow on this work. First, the authors did not suggest how the process could have started in a prebiotic world—how the first version of the ribozyme (composed of about 76 ribonucleotides) could have been produced. Second, the molecules that the ribozyme operated upon (i.e., the substrates) included preprepared, fully formed ribozymes split into two halves and the exact complement of the fully formed ribozyme split into two halves. In other words, the solution contained one complete ribozyme and many copies of each of four half-ribozymes. The complete ribozyme first created a single bond joining halves 1 and 2, producing a complementary copy of itself. The complementary ribozyme then acted to create a single bond joining halves 3 and 4, producing a copy of the original ribozyme. Some other processes had to produce the four half-ribozymes with high purity. A third limitation comes from the observation that this process of replication cannot experience chemical evolution: it only combines two preexisting substrates. For chemical evolution to occur, errors in replication (i.e., mutations) must lead to a selective advantage, such as faster reproduction (a type of natural selection), and be passed to their progeny so that improvements will dominate the population over time (“survival of the fittest”). In the process of RNA replication described in this paper, variations in the components could have resulted in an improved ribozyme, but the improved ribozyme could not pass its improvement on to the next generation (the next two half-ribozymes that it joins). The improved ribozyme, like its predecessors, simply combines two components, not necessarily creating copies of its new and improved self. This is because the supply of components cannot “evolve” themselves because they cannot receive feedback (through natural selection) about which versions of the components are preferred.
The very origin of the first organisms presents at least an appearance of a paradox because a certain minimum level of complexity is required to make self-replication possible at all; high-fidelity replication requires additional functionalities that need even more information to be encoded (Penny, 2005).
The crucial question in the study of the origin of life is how the Darwin-Eigen cycle started—how was the minimum complexity that is required to achieve the minimally acceptable replication fidelity attained? In even the simplest modern systems, such as RNA viruses with the replication fidelity of only about 10^3 and viroids that replicate with the lowest fidelity among the known replicons (about 10^2; Gago, et al., 2009), replication is catalyzed by complex protein polymerases. The replicase itself is produced by translation of the respective mRNA(s), which is mediated by the immensely complex ribosomal apparatus. Hence, the dramatic paradox of the origin of life is that, to attain the minimum complexity required for a biological system to start on the Darwin-Eigen spiral, a system of a far greater complexity appears to be required. How such a system could evolve is a puzzle that defeats conventional evolutionary thinking, all of which is about biological systems moving along the spiral; the solution is bound to be unusual.
Eugene V. Koonin: The Logic of Chance: The Nature and Origin of Biological Evolution
The primary incentive behind the theory of self-replicating systems that Manfred Eigen outlined was to develop a simple model explaining the origin of biological information and, hence, of life itself. Eigen’s theory revealed the existence of the fundamental limit on the fidelity of replication (the Eigen threshold): If the product of the error (mutation) rate and the information capacity (genome size) is below the Eigen threshold, there will be stable inheritance and hence evolution; however, if it is above the threshold, the mutational meltdown and extinction become inevitable (Eigen, 1971). The Eigen threshold lies somewhere between 1 and 10 mutations per round of replication (Tejero, et al., 2011); regardless of the exact value, staying above the threshold fidelity is required for sustainable replication and so is a prerequisite for the start of biological evolution
(see Figure 12-1A).
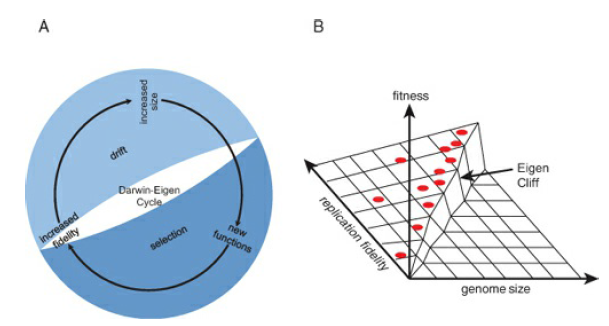
Indeed, the very origin of the first organisms presents at least an appearance of a paradox because a certain minimum level of complexity is required to make self-replication possible at all; high-fidelity replication requires additional functionalities that need even more information to be encoded (Penny, 2005). However, the replication fidelity at a given point in time limits the amount of information that can be encoded in the genome. What turns this seemingly vicious circle into the (seemingly) unending spiral of increasing complexity—the Darwin-Eigen cycle, following the terminology introduced by David Penny (Penny, 2005)—is a combination of natural selection with genetic drift. Even small gains in replication fidelity are advantageous to the system, if only because of the decrease of the reproduction cost as a result of the increasing yield of viable copies of the genome. In itself, a larger genome is more of a liability than an advantage because of higher replication costs. However, moderate genome increase, such as by duplication of parts of the genome or by recombination, can be fixed via genetic drift in small populations. Replicators with a sufficiently high fidelity can take advantage of such randomly fixed and initially useless genetic material by evolving new functions, without falling off the “Eigen cliff” (see Figure 12-1B). Among such newly evolved, fitness-increasing functions will be those that increase replication fidelity, which, in turn, allows a further increase in the amount of encoded information. And so the Darwin- Eigen cycle recapitulates itself in a spiral progression, leading to a steady increase in genome complexity (see Figure 12-1A). The crucial question in the study of the origin of life is how the Darwin-Eigen cycle started—how was the minimum complexity that is required to achieve the minimally acceptable replication fidelity attained? In even the simplest modern systems, such as RNA viruses with the replication fidelity of only about 10^3 and viroids that replicate with the lowest fidelity among the known replicons (about 10^2; Gago, et al., 2009), replication is catalyzed by complex protein polymerases. The replicase itself is produced by translation of the respective mRNA(s), which is mediated by the immensely complex ribosomal apparatus. Hence, the dramatic paradox of the origin of life is that, to attain the minimum complexity required for a biological system to start on the Darwin-Eigen spiral, a system of a far greater complexity appears to be required. How such a system could evolve is a puzzle that defeats conventional evolutionary thinking, all of which is about biological systems moving along the spiral; the solution is bound to be unusual.
My comment: Or the solution might be outside the realm of philosophical naturalism, that is, intelligent design ?!!
In the next sections, we first examine the potential of a top-down approach based on the analysis of extant genes, to obtain clues on possible origins of replicator systems. We then discuss the bottom-up approach. The case for a complex RNA World from protein domain evolution: The top-down view As pointed out earlier, the translation system is the only complex ensemble of genes that is conserved in all extant cellular life forms. With about 60 protein-coding genes and some 40 structural RNA genes universally conserved, the modern translation system is the best-preserved relic of the LUCA(S) and the strongest available piece of evidence that some form of LUCA(S) actually existed . Given this extraordinary conservation of the translation system, comparison of orthologous sequences reveals very little, if anything, about its origins: The emergence of the
translation system is beyond the horizon of the comparison of extant life forms. Indeed, comparative genomic reconstructions of the gene repertoire of LUCA(S) point to a complex translation system that includes at least 18 of the 20 aminoacyl-tRNA synthetases (aaRS), several translation factors, at least 40 ribosomal proteins, and several enzymes involved in rRNA and tRNA modification. It appears that the core of the translation system was already fully shaped in LUCA(S) (Anantharaman, et al., 2002). Fortunately, sequence and structure comparisons of protein and RNA components within the translation system itself are informative, thanks to the extensive paralogy among the respective genes. Whenever a pair of paralogous genes is assigned to LUCA(S), the respective duplication must have been a more ancient event, so reconstruction of the series of ancient duplications opens a window into very early stages of evolution. The story of the paralogous aaRS is particularly revealing. The aaRS form two distinct classes of ten specificities each (that is, each class is responsible for the recognition and activation of ten amino acids), with unrelated catalytic domains and distinct sets of accessory domains. The catalytic domains of the Class I and Class II aaRS belong to the Rossmann fold and the biotin synthase fold, respectively. The analysis of the evolutionary histories of these protein folds has far-reaching implications for the early evolution of the translation system and beyond (Aravind, et al., 2002). The catalytic domains of Class I aaRS form but a small twig in the evolutionary tree of the Rossmann fold domains (see Figure 12-2A).

Figure 12-2A
Diversification of protein domains, crystallization of the translation system, and the LUCA(S): Evolution of the Rossmann fold–like nucleotide-binding domains. Based on data from Aravind, et al., 2002. Only the better-known proteins are indicated. USPA = Universal stress protein A; ETFP = electron transfer flavoprotein; vWA = Von Willebrand A factor; Toprim = catalytic domain of topoisomerases, primases, and some nucleases; Receiver = a component of prokaryotic two-component signaling systems; TIR = a widespread protein-protein interaction domain in prokaryotic and eukaryotic signaling systems; Sir2 = protein (in particular, histone) deacetylase; Methylase = diverse methyltransferases. For details, see (Aravind, et al., 2002) and references therein.
Thus, the appearance of the common ancestor of the aaRS is preceded by a number of nodes along the evolutionary path from the primitive, ancestral domain to the highly diversified state that corresponds to LUCA(S). So a substantial diversity of Rossmann fold domains evolved prior to the series of duplications that led to the emergence of the aaRS of different specificities, which itself antedates LUCA(S) (see Figure 12-2A). A similar evolutionary pattern is implied by the analysis of the biotin synthase domain that gave
rise to Class II aaRS. Thus, even within these two folds alone, remarkable structural and functional complexity of protein domains had evolved before the full-fledged RNA-protein machinery of translation resembling the modern system was in place.
Ribozymes and the RNA World
The Central Dogma of molecular biology (Crick, 1970) states that, in biological systems, information is transferred from DNA to protein through an RNA intermediate (Francis Crick added the possibility of reverse information flow from RNA to DNA after the discovery of RT): DNA
experiments, ribozymes have been evolved to catalyze an extremely broad variety of reactions. Box 12-1 lists some of the most biologically relevant ribozyme-catalyzed reactions.

Notably, all three elementary reactions that are required for translation—namely (i) amino acid activation through the formation of aminoacyl-AMP, (ii) (t)RNA aminoacylation, and (iii) transpeptidation (the peptidyltransferase reaction)—have been successfully modeled with ribozymes. The selfaminoacylation reaction that is key to the origin of the primordial RNA-only adaptors (the RNA analog of aaRS) has been selected in vitro with relative ease. Strikingly, the best of the resulting ribozymes catalyze this reaction with a rate and specificity greater than those of the respective aaRS, and very short oligonucleotides possessing this activity have been selected (Turk, et al., 2010). Understandably, major effort has focused on the demonstration of nucleotide polymerization and, ultimately, RNA replication catalyzed by ribozymes, the central processes in the hypothetical primordial RNA World. The outcome of the experiments aimed at the creation of ribozyme replicases so far has been mixed (Cheng and Unrau, 2010). Ribozymes have been obtained that are capable of extending a primer annealed to a template (Johnston, et al., 2001); initially, the ribozymes with this activity could function only by specific base-pairing to the template, but subsequently general ribozyme polymerases of this class have been evolved through additional selection (Lincoln and Joyce, 2009). The latest breakthrough in the field of polymerase ribozymes has been published at the time of the final editing of this chapter: an active endonuclease ribozyme was produced using a
ribozyme polymerase that itself was constructed by recombining two pre-existing ribozymes, potentially, a plausible route for pre-biological evolution (Wochner, et al., 2011). All this progress notwithstanding, the ribozyme polymerases that are currently available are a far cry from processive, sufficiently accurate (in terms of the Eigen threshold) replicases, capable of catalyzing the replication of exogenous templates and themselves. Enzymes with such properties appear to be a conditio sine qua non for the evolution of the hypothetical RNA World. Besides, even the available ribozymes with the limited RNA polymerase capacity are rather complex molecules that consist of some 200 nucleotides and could be nontrivial to evolve in the prebiotic setting.
Is the evidence in support of any of these models and scenarios compelling? Of course, the question already implies a negative answer. We do have some strong hints, even if these are a far cry from a coherent scenario of the earliest stages of evolution of biological information transmission. First, consider the apparent logical inevitability of an RNA World: What other starting point for the evolution of the translation system could there be? Second, comparative analysis of the translation system components does point to a much greater role of RNA in ancestral translation, compared to the modern system—notably, the decisive function of RNA as the determinant of amino acid–codon specificity. Third, ribozymes are impressive (if in general far inferior to proteins) in their catalytic versatility and efficiency. Thirty years ago, no catalytic activity was reported for any RNA molecule to catalyze any reaction at all; now we are aware of dozens of ribozyme activities, including some, such as highly efficient aminoacylation, that get the translation system going. However, this is about all the good news; the rest is more like a sobering cold shower. For all the advances of “ribozymology,” no ribozyme polymerase comes close to what is required if we are to accept an RNA-only replicator system as a key intermediate stage in the evolution of life. Nor are any ribozymes capable of catalyzing the synthesis of nucleotides or even their sugar moieties. Even sweeping all these problems under the proverbial rug, the path from a putative RNA World to the translation system is incredibly steep. The general idea of a function(s) for abiogenic amino acids and possibly peptides in the RNA World, such as the role of ribozyme cofactors (see the discussion in the preceding sections), appears fruitful and is compatible with experimental data. Nevertheless, breaking the evolution of the translation system into incremental steps, each associated with a biologically plausible selective advantage, is extremely difficult even within a speculative scheme let alone experimentally. The triplicase/protoribosome hypothesis is attractive as an attempt to explain the origin of translation and replication in one sweep, but is this scenario realistic? The triplicase itself would have to be an extremely complex, elaborate molecular machine, leaving one with the suspicion that, all its attraction notwithstanding, the triplicase might not be the most likely solution to the origin of translation problem.
The overall situation in the origin of life field appears rather grim. Even under the (highly nontrivial) assumption that monomers such as NTP are readily available, the problem of the synthesis of sufficiently stable, structurally regular polymers (RNA) is formidable, and the origin of replication and translation from such primordial RNA molecules could be an even harder problem. As emphasized repeatedly in this book, evolution by natural selection and drift can begin only after replication with sufficient fidelity is established. Even at that stage, the evolution of translation remains highly problematic. The emergence of the first replicator system, which represented the “Darwinian breakthrough,” was inevitably preceded by a succession of complex, difficult steps for which biological evolutionary mechanisms were not accessible (see Figure 12-6).
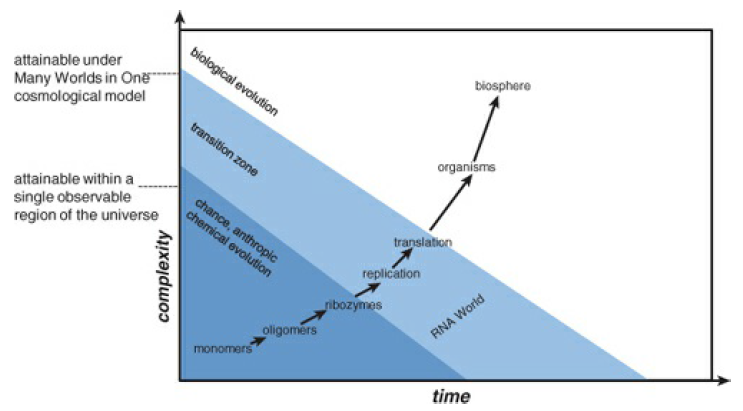
Figure 12-6 The prebiological and biological stages of the origin of life: the transition from anthropic causality to biological evolution.
Even considering environments that could facilitate these processes, such as networks of inorganic compartments at hydrothermal vents, multiplication of the probabilities for these steps could make the emergence of the first replicators staggeringly improbable.
The ultimate enigma of the origin of life
The origin of life—or, to be more precise, the origin of the first replicator systems and the origin of translation—remains a huge enigma, and progress in solving these problems has been very modest—in the case of translation, nearly negligible. Some potentially fruitful observations and ideas exist, such as the discovery of plausible hatcheries for life, the networks of inorganic compartments at hydrothermal vents, and the chemical versatility of ribozymes that fuels the RNA World hypothesis. However, these advances remain only preliminaries, even if important ones, because they do not even come close to a coherent scenario for prebiological evolution, from the first organic molecules to the first replicator systems, and from these to bona fide biological entities in which information storage and function are partitioned between distinct classes of molecules (nucleic acids and proteins, respectively). In my view, all advances notwithstanding, evolutionary biology is and will remain woefully incomplete until there is at least a plausible, even if not compelling, origin of life scenario.
Under this replication-centered perspective, the emergence of complexity is an enigma: Why are there numerous life forms that are far more complex than the minimal, simplest device for replication? We cannot know “for sure” what these minimally complex devices are, but there are excellent candidates —namely, the simplest autotrophic bacteria and archaea, such as Pelagibacter ubique or Prochlorococcus sp. These organisms get by with about 1,300 genes without using any organic molecules, and generally without any dependence on other life forms. Incidentally, these are also the most “successful” organisms on Earth. They have the largest populations that have evolved under the strongest selection pressure—and consequently have the most “streamlined” genomes. A complete biosphere consisting of such highly effective unicellular organisms is easily imaginable; indeed, the Earth biota prior to the emergence of eukaryotes (that is, probably for the 2 billion years of the evolution of life or so) must have resembled this image much closer than today’s biosphere (although more complex prokaryotes certainly existed even at that time).
So why complex organisms?
One answer that probably appeared most intuitive to biologists and to everyone else interested in evolution over the centuries is that the more complex organisms are also the more fit. This view is demonstrably false. Indeed, to accentuate the paradox of complexity, the general rule is the opposite: The more complex a life form is, the smaller effective population size it has, and so the less successful it is, under the only sensible definition of evolutionary success. This pattern immediately suggests that the answer to the puzzle of complexity emergence could be startlingly simple: Just turn this trend around and posit that the smaller the effective population size, the weaker the selection intensity, hence the greater the chance of non-adaptive evolution of complexity. This is indeed the essence of the population-genetic non-adaptive concept that Lynch propounded.
All this progress notwithstanding, the ribozyme polymerases that are currently available are a far cry from processive, sufficiently accurate (in terms of the Eigen threshold) replicases, capable of catalyzing the replication of exogenous templates and themselves. Thirty years ago, no catalytic activity was reported for any RNA molecule to catalyze any reaction at all; now we are aware of dozens of ribozyme activities, including some, such as highly efficient aminoacylation, that get the translation system going. However, this is about all the good news; the rest is more like a sobering cold shower.
The origin of life—or, to be more precise, the origin of the first replicator systems and the origin of translation—remains a huge enigma, and progress in solving these problems has been very modest—in the case of translation, nearly negligible.
https://www.liebertpub.com/doi/10.1089/ast.2022.0027
Last edited by Otangelo on Mon Sep 26, 2022 10:50 am; edited 24 times in total

