https://reasonandscience.catsboard.com/t2221-the-hardware-and-software-of-the-cell-evidence-of-design
Paul Davies: the fifth miracle page 62
Due to the organizational structure of systems capable of processing algorithmic (instructional) information, it is not at all clear that a monomolecular system – where a single polymer plays the role of catalyst and informational carrier – is even logically consistent with the organization of information flow in living systems, because there is no possibility of separating information storage from information processing (that being such a distinctive feature of modern life). As such, digital–first systems (as currently posed) represent a rather trivial form of information processing that fails to capture the logical structure of life as we know it. 1
We need to explain the origin of both the hardware and software aspects of life, or the job is only half finished. Explaining the chemical substrate of life and claiming it as a solution to life’s origin is like pointing to silicon and copper as an explanation for the goings-on inside a computer. It is this transition where one should expect to see a chemical system literally take-on “a life of its own”, characterized by informational dynamics which become decoupled from the dictates of local chemistry alone (while of course remaining fully consistent with those dictates). Thus the famed chicken-or-egg problem (a solely hardware issue) is not the true sticking point. Rather, the puzzle lies with something fundamentally different, a problem of causal organization having to do with the separation of informational and mechanical aspects into parallel causal narratives. The real challenge of life’s origin is thus to explain how instructional information control systems emerge naturally and spontaneously from mere molecular dynamics.
Software and hardware are irreducible complex and interdependent. There is no reason for information processing machinery to exist without the software, and vice versa.
Systems of interconnected software and hardware are irreducibly complex. 2
Is the cell really a machine?
Following the Second World War, the pioneering ideas of cy- bernetics, information theory, and computer science captured the imagination of biologists, providing a new vision of the MCC that was translated into a highly successful experimental research program, which came to be known as ‘molecular biology’ ( Keller, 1995; Morange, 1998; Kay, 20 0 0 ). At its core was the idea of the computer, which, by introducing the conceptual distinction between ‘software’ and ‘hardware’, directed the attention of re- searchers to the nature and coding of the genetic instructions (the software) and to the mechanisms by which these are im- plemented by the cell’s macromolecular components (the hardware).
https://sci-hub.ren/10.1016/j.jtbi.2019.06.002
All cellular functions are irreducibly complex 3
http://reasonandscience.heavenforum.org/t2179-the-cell-is-a-interdependent-irreducible-complex-system
chemist Wilhelm Huck, professor at Radboud University Nijmegen 5
A working cell is more than the sum of its parts. "A functioning cell must be entirely correct at once, in all its complexity,"
Paul Davies, the fifth miracle page 53:
Pluck the DNA from a living cell and it would be stranded, unable to carry out its familiar role. Only within the context of a highly specific molecular milieu will a given molecule play its role in life. To function properly, DNA must be part of a large team, with each molecule executing its assigned task alongside the others in a cooperative manner. Acknowledging the interdependability of the component molecules within a living organism immediately presents us with a stark philosophical puzzle. If everything needs everything else, how did the community of molecules ever arise in the first place? Since most large molecules needed for life are produced only by living organisms, and are not found outside the cell, how did they come to exist originally, without the help of a meddling scientist? Could we seriously expect a Miller-Urey type of soup to make them all at once, given the hit-and-miss nature of its chemistry?
Being part of a large team,cooperative manner,interdependability,everything needs everything else, are just other words for irreducibility and interdependence.
For a nonliving system, questions about irreducible complexity are even more challenging for a totally natural non-design scenario, because natural selection — which is the main mechanism of Darwinian evolution — cannot exist until a system can reproduce. For an origin of life we can think about the minimal complexity that would be required for reproduction and other basic life-functions. Most scientists think this would require hundreds of biomolecular parts, not just the five parts in a simple mousetrap or in my imaginary LMNOP system. And current science has no plausible theories to explain how the minimal complexity required for life (and the beginning of biological natural selection) could have been produced by natural process before the beginning of biological natural selection.
If we attempt to build model systems for studying chemical evolution prior to the time when information-bearing templates were part of the developing prebiotic system, we immediately encounter three problems: (1) very few experimental systems have been developed for such studies; (2) a complex chemical system has no obvious repository for the storage of useful chemical information; (3) the rules governing the evolution of purely chemical systems are not known. 4
Perry Marshall, Evolution 2.0, page 153:
Wanna Build a Cell? A DVD Player Might Be Easier Imagine that you’re building the world’s first DVD player. What must you have before you can turn it on and watch a movie for the first time?
A DVD. How do you get a DVD? You need a DVD recorder first. How do you make a DVD recorder? First you have to define the language. When Russell Kirsch (who we met in chapter ) created the world’s first digital image, he had to define a language for images first. Likewise you have to define the language that gets written on the DVD, then build hardware that speaks that language. Language must be defined first. Our DVD recorder/player problem is an encoding-decoding problem, just like the information in DNA. You’ll recall that communication, by definition, requires four things to exist:
1. A code
2. An encoder that obeys the rules of a code
3. A message that obeys the rules of the code
4. A decoder that obeys the rules of the code
These four things—language, transmitter of language, message, and receiver of language—all have to be precisely defined in advance before any form of communication can be possible at all.
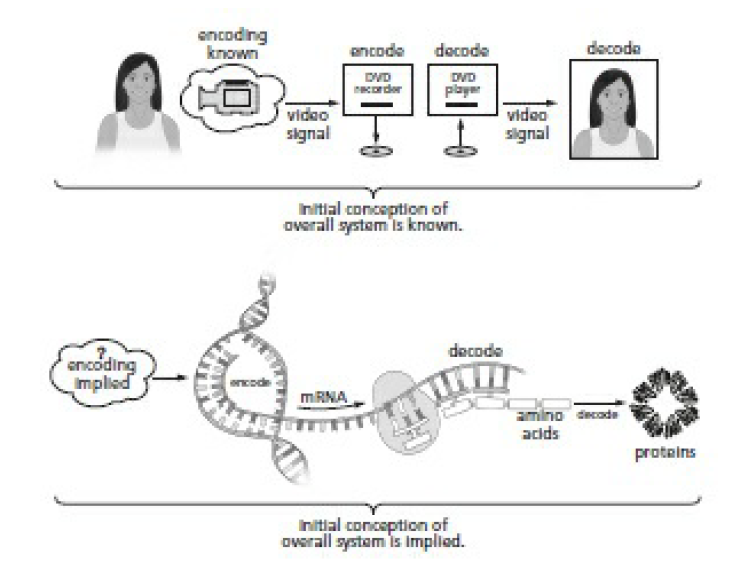
A camera sends a signal to a DVD recorder, which records a DVD. The DVD player reads the DVD and converts it to a TV signal. This is conceptually identical to DNA translation. The only difference is that we don’t know how the original signal—the pattern in the first DNA strand—was encoded. The first DNA strand had to contain a plan to build something, and that plan had to get there somehow. An original encoder that translates the idea of an organism into instructions to build the organism (analogous to the camera) is directly implied.
The rules of any communication system are always defined in advance by a process of deliberate choices. There must be prearranged agreement between sender and receiver, otherwise communication is impossible. By definition, a communication system cannot evolve from something simpler because evolution itself requires communication to exist first. You can’t make copies of a message without the message, and you can’t create a message without first having a language. And before that, you need intent. A code is an abstract, immaterial, nonphysical set of rules. There is no physical law that says ink on a piece of paper formed in the shape T-R-E-E should correspond to that large leafy organism in your front yard. You cannot derive the local rules of a code from the laws of physics, because hard physical laws necessarily exclude choice. On the other hand, the coder decides whether “1” means “on” or “off.” She decides whether “0” means “off” or “on.” Codes, by definition, are freely chosen. The rules of the code come before all else. These rules of any language are chosen with a goal in mind: communication, which is always driven by intent. That being said, conscious beings can evolve a simple code into a more complex code—if they can communicate in the first place. But even simple grunts and hand motions between two humans who share no language still require communication to occur. Pointing to a table and making a sound that means “table” still requires someone to recognize what your pointing finger means.
https://reasonandscience.catsboard.com/t2221-the-hardware-and-software-of-the-cell-evidence-of-design
In order to explain the origin of life, the origin of the physical parts, that is DNA, RNA, organelles , proteins, enzymes etc. of the cell must be explained, and the origin of the information and various code systems of the cell. Following excerpt will elucidate why the origin of both, the software, and the hardware are best explained through the action of a intentional creator.
Replication upon which mutations and natural selection act could not begin prior when life started and cell's began with self-replication. According to geneticist Michael Denton, the break between the nonliving and the living world ‘represents the most dramatic and fundamental of all the discontinuities of nature. Before that remarkable event, a fully operating cell had to be in place, various organelles, enzymes, proteins, DNA, RNA, tRNA, mRNA, and a a extraordinarily complex machinery, that is : a complete DNA replication machinery, topoisomerases for replication and chromosome segregation functions, a DNA repair system, RNA polymerase and transcription factors, a fully operational ribosome for translation, including 20 aminoacyl-tRNA synthases, tRNA and the complex machinery to synthesize them, proteins for posttranslational modifications and chaperones for correct folding of a series of essential proteins, FtsZ microfilaments for celldivision and formation of cell shape, a transport system for proteins etc., a complex metabolic system consistent of several different enzymes for energy generation, lipid biosynthesis to make the cell membrane, and machinery for nucleosynthesis.
This constitutes a minimal set of basic parts, superficially described. If one, just ONE of these parts is missing, the cell will not operate. That constitutes a interdependent , interlocked and irreducibly complex biological system of extraordinary complexity, which had to arise ALL AT ONCE. No step by step build up over a long period of time is possible.
http://reasonandscience.heavenforum.org/t2110-what-might-be-a-protocells-minimal-requirement-of-parts#3797
That constitutes a formidable catch22 problem.
Biochemist David Goodsell describes the problem, "The key molecular process that makes modern life possible is protein synthesis, since proteins are used in nearly every aspect of living. The synthesis of proteins requires a tightly integrated sequence of reactions, most of which are themselves performed by proteins." and continues : this "is one of the unanswered riddles of biochemistry: which came first, proteins or protein synthesis? If proteins are needed to make proteins, how did the whole thing get started?"45 The end result of protein synthesis is required before it can begin. To make it more clear what we are talking about:
That chain is constituted by INITIATION OF TRANSCRIPTION, CAPPING, ELONGATION, SPLICING, CLEAVAGE,POLYADENYLATION AND TERMINATION, EXPORT FROM THE NUCLEUS TO THE CYTOSOL, INITIATION OF PROTEIN SYNTHESIS (TRANSLATION), COMPLETION OF PROTEIN SYNTHESIS AND PROTEIN FOLDING. In order for evolution to work, the robot-like working machinery and assembly line must be in place, fully operational.
Jacques Monod noted: "The code is meaningless unless translated. The modern cell's translating machinery consists of at least fifty macromolecular components which are themselves coded in DNA: the code cannot be translated otherwise than by products of translation." (Scientists now know that translation actually requires more than a hundred proteins.)
http://reasonandscience.heavenforum.org/t2059-catch22-chicken-and-egg-problems?highlight=catch22
Furthermore, to build up this system, following conditions in a primordial earth would have to be met :
( Agents Under Fire: Materialism and the Rationality of Science, pgs. 104-105 (Rowman & Littlefield, 2004). HT: ENV.)
Availability. Among the parts available for recruitment to form the system, there would need to be ones capable of performing the highly specialized tasks of individual parts, even though all of these items serve some other function or no function.
Synchronization. The availability of these parts would have to be synchronized so that at some point, either individually or in combination, they are all available at the same time.
Localization. The selected parts must all be made available at the same ‘construction site,’ perhaps not simultaneously but certainly at the time they are needed.
Coordination. The parts must be coordinated in just the right way: even if all of the parts of a system are available at the right time, it is clear that the majority of ways of assembling them will be non-functional or irrelevant.
Interface compatibility. The parts must be mutually compatible, that is, ‘well-matched’ and capable of properly ‘interacting’: even if sub systems or parts are put together in the right order, they also need to interface correctly.
http://reasonandscience.heavenforum.org/t1468-irreducible-complexity#2133
Fred Hoyles example is not far fetched but rather a excellent illustration. If it is ridiculous to think that a perfectly operational 747 Jumbo-jet could come into existence via a lucky accident, then it is likewise just as illogical to think that such a sophisticated organism like a first living cell could assemble by chance . It gets even more absurd to think that the OOL would also form by chance and have the capability to reproduce. Life cannot come from non-life even if given infinite time and oportunities. If life could spontaneously arise from non-life than it still should be doing that today. Hence the Law of Biogenesis: The principle stating that life arises from pre-existing life, not from non-living matter. Life is clearly best explained through the wilful action of a extraordinarily intelligent and powerful designer.
http://reasonandscience.heavenforum.org/t1279p30-abiogenesis-is-impossible#4171
that is the hardware part, which you can compare to a computer with hard disk, cabinet, cpu etc.
The software
BIOLOGICAL CELL AND ITS SYSTEM SOFTWARE 3
Abstract: Looking for a role that DNA has in the cell, we are elaborating that DNA can be viewed as a Cell Operating System. In the paper we consider several aspects of that approach and we propose some possible explanations for some of the not yet understood phenomena in molecular genetics in terms of systems software.
We recognize the operon as a monitor a piece of software that contains the data, but also it controls the access over its data. It is an object in a sense that it contains data structures (genes) and control structures (operators) for accessing the data. This suggests that the prokaryote systems software is object oriented. In the cell hierarchical control system, the operon is the smallest object. The exons can be viewed as subprograms, programs, or data of modular software. In some cases they are pieces of reusable software, since combination of exons can produce different programs, as in the case with program preparation for antibodies.
Secondly, you need coded, specified, complex information. Thats the software. And constitutes the second major hurdle that buries any naturalistic just so fairy tale stories and fantasies. In following paper :
Origin and evolution of the genetic code: the universal enigma
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3293468/
In our opinion, despite extensive and, in many cases, elaborate attempts to model code optimization, ingenious theorizing along the lines of the coevolution theory, and considerable experimentation, very little definitive progress has been made.
Summarizing the state of the art in the study of the code evolution, we cannot escape considerable skepticism. It seems that the two-pronged fundamental question: “why is the genetic code the way it is and how did it come to be?”, that was asked over 50 years ago, at the dawn of molecular biology, might remain pertinent even in another 50 years. Our consolation is that we cannot think of a more fundamental problem in biology.
http://reasonandscience.heavenforum.org/t2001-origin-and-evolution-of-the-genetic-code-the-universal-enigma
The genetic code is nearly optimal for allowing additional information within protein-coding sequences
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1832087/
DNA sequences that code for proteins need to convey, in addition to the protein-coding information, several different signals at the same time. These “parallel codes” include binding sequences for regulatory and structural proteins, signals for splicing, and RNA secondary structure. Here, we show that the universal genetic code can efficiently carry arbitrary parallel codes much better than the vast majority of other possible genetic codes. This property is related to the identity of the stop codons. We find that the ability to support parallel codes is strongly tied to another useful property of the genetic code—minimization of the effects of frame-shift translation errors. Whereas many of the known regulatory codes reside in nontranslated regions of the genome, the present findings suggest that protein-coding regions can readily carry abundant additional information.
if we employ weightings to allow for biases in translation, then only 1 in every million random alternative codes generated is more efficient than the natural code. We thus conclude not only that the natural genetic code is extremely efficient at minimizing the effects of errors, but also that its structure reflects biases in these errors, as might be expected were the code the product of selection.
Fazale Rana wrote in his book Cell's design: In 1968, Nobel laureate Francis Crick argued that the genetic code could not undergo significant evolution. His rationale is easy to understand. Any change in codon assignments would lead to changes in amino acids in every polypeptide made by the cell. This wholesale change in polypeptide sequences would result in a large number of defective proteins. Nearly any conceivable change to the genetic code would be lethal to the cell.
Even if the genetic code could change over time to yield a set of rules that allowed for the best possible error-minimization capacity, is there enough time for this process to occur? Biophysicist Hubert Yockey addressed this question. He determined that natural selection would have to explore 1.40 x 10^70 different genetic codes to discover the universal genetic code found in nature. The maximum time available for it to originate was estimated at 6.3 x 10^15 seconds. Natural selection would have to evaluate roughly 10^55 codes per second to find the one that's universal. Put simply, natural selection lacks the time necessary to find the universal genetic code.
http://reasonandscience.heavenforum.org/t1404-the-genetic-code-is-nearly-optimal-for-allowing-additional-information-within-protein-coding-sequences
The cell converts the information carried in an mRNA molecule into a protein molecule. This feat of translation was a focus of attention of biologists in the late 1950s, when it was posed as the “coding problem”: how is the information in a linear sequence of nucleotides in RNA translated into the linear sequence of a chemically quite different set of units—the amino acids in proteins?
The first scientist after Watson and Crick to find a solution of the coding problem, that is the relationship between the DNA structure and protein synthesis was Russian physicist George Gamow. Gamow published in the October 1953 issue of Nature a solution called the “diamond code”, an overlapping triplet code based on a combinatorial scheme in which 4 nucleotides arranged 3-at-a-time would specify 20 amino acids. Somewhat like a language, this highly restrictive code was primarily hypothetical, based on then-current knowledge of the behavior of nucleic acids and proteins.
The concept of coding applied to genetic specificity was somewhat misleading, as translation between the four nucleic acid bases and the 20 amino acids would obey the rules of a cipher instead of a code. As Crick acknowledged years later, in linguistic analysis, ciphers generally operate on units of regular length (as in the triplet DNA scheme), whereas codes operate on units of variable length (e.g., words, phrases). But the code metaphor worked well, even though it was literally inaccurate, and in Crick’s words, “‘Genetic code’ sounds a lot more intriguing than ‘genetic cipher’.”
Question: how did the translation of the triplet anti codon to amino acids, and its assignment, arise ? There is no physical affinity between the anti codon and the amino acids. What must be explained, is the arrangement of the codon " words " in the standard codon table which is highly non-random, redundant and optimal, and serves to translate the information into the amino acid sequence to make proteins, and the origin of the assignment of the 64 triplet codons to the 20 amino acids. That is, the origin of its translation. The origin of a alphabet through the triplet codons is one thing, but on top, it has to be translated to a other " alphabet " constituted through the 20 amino acids. That is as to explain the origin of capability to translate the english language into chinese. We have to constitute the english and chinese language and symbols first, in order to know its equivalence. That is a mental process.
http://reasonandscience.heavenforum.org/t2057-origin-of-translation-of-the-4-nucleic-acid-bases-and-the-20-amino-acids-and-the-universal-assignment-of-codons-to-amino-acids
1) http://arxiv.org/pdf/1207.4803v2.pdf
2) http://www.evolutionnews.org/2012/07/software_machin062211.html
3) http://ciit.finki.ukim.mk/data/papers/2CiiT/2CiiT-19.pdf
4) Origins of Life on the Earth and in the Cosmos page 185
Last edited by Otangelo on Tue Jul 16, 2024 1:41 pm; edited 13 times in total

