Catch22, chicken and egg problems in biology and biochemistry
https://reasonandscience.catsboard.com/t2059-catch22-chicken-and-egg-problems-in-biology-and-biochemistry

Life’s First Molecule Was Protein, Not RNA, New Model Suggests
For scientists studying the origin of life, one of the greatest chicken-or-the-egg questions is: Which came first — proteins or nucleic acids like DNA and RNA? 24
Information is required to extract energy. But extract energy is required to store information. What came first?
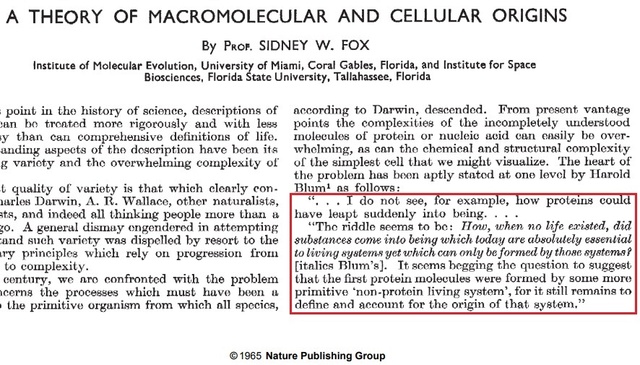
The Catch22 riddle in 1965
https://sci-hub.ren/https://www.nature.com/articles/205328a0
what changed, is that rather than closing the gaps, they have become wider and wider, and naturalistic proposals more and more unrealistic!
In Joseph Heller’s classic novel about World War II,Catch-22, an aviator could be excused from combat duty for being crazy. But a rule specified that he first had to request an excuse, and anyone who requested an excuse from combat duty was obviously not crazy, so such requests were invariably denied. The rule that made it impossible to be excused from combat duty was called “Catch-22.”

Oxygen and life have a catch 22 relationship
Catch 22 is a situation in which an action has consequences which make impossible to pursue that action. Oxygen is very harmful to life. At the same time oxygen is needed to provide the ozone layer which protects life from ultraviolet radiation (UVR) coming from the sun. If Cyanobacteria came before oxygen, because it is the cause of oxygen, then Cyanobacteria would have had to develop several forms of protection to mitigate the damage from UVR: avoidance, scavenging, screening, repair, and programmed cell death. However, UVR damage is immediate and the time needed to “evolve” protection against it via natural selection, incredibly slow. So, UVR damage would occur before any such defense mechanisms could evolve. 3
Iron-sulfur clusters
Sulfur is an essential element, being a constituent of many proteins and cofactors. Iron-sulfur (FeS) centers are essential protein cofactors in all forms of life. Various biosynthetic pathways were found to be tightly interconnected through complex crosstalk mechanisms that crucially depend on the bio-availability of the metal ions iron, molybdenum, tungsten, nickel, copper, and zinc. Proteins requiring Fe/S clusters in their active site have been localized in mitochondria, cytosol, and nucleus where they are involved in rather diverse functions such as the TCA cycle, amino acid biosynthesis, bacterial and mitochondrial respiration, co-factor biosynthesis, ribosome assembly, regulation of protein translation, DNA replication and DNA repair. Hence the process of iron-sulfur biosynthesis is essential to almost all forms of life. The prevalence of these proteins on the metabolic pathways of most organisms leads some scientists to theorize that iron-sulfur compounds had a significant role in the origin of life in the iron-sulfur world theory. The iron-sulfur world hypothesis is a set of proposals for the origin of life and the early evolution of life advanced in a series of articles between 1988 and 1992 by Günter Wächtershäuser. FeS cluster assembly is a complex process involving the mobilization of Fe and S atoms from storage sources, their assembly into [Fe-S] form, their transport to specific cellular locations, and their transfer to recipient apoproteins. Nar1 is an essential component of a cytosolic Fe/S protein assembly machinery. Required for maturation of extramitochondrial Fe/S proteins. 12 Thus, Nar1 is both a target and a component of the cellular Fe/S protein biogenesis machinery creating an interesting “chicken and egg” situation for its maturation process Conserved Iron–Sulfur (Fe–S) clusters are found in a growing family of metalloproteins that are implicated in prokaryotic and eukaryotic DNA replication and repair. 20
Therefore, they had to exist prior to life beginning, since DNA replication enzymes and proteins depend on them. They require however also complex proteins and enzymes to be synthesized. That's a classical chicken/egg problem.
Ribonucleotide reductase
This is one of the most essential enzymes of life. Ribonucleotide reduction is the only pathway for de novo synthesis of deoxyribonucleotides in extant organisms. This chemically demanding reaction is catalyzed by ribonucleotide reductase (RNR). The mechanism has been deemed unlikely to be catalyzed by a ribozyme, creating an enigma regarding how the building blocks for DNA were synthesized at the transition from RNA to DNA-encoded genomes. Biosynthesis DNA is made from RNA. The deoxynucleotides are made from nucleotides with ribonucleotide reductases (RNRs), producing uracil-DNA or u-DNA. The uracil is then converted to thymine by adding a methyl group, making thymine-DNA or t-DNA, the kind that is actually used. The reaction catalyzed by RNR is strictly conserved in all living organisms. Furthermore, RNR plays a critical role in regulating the total rate of DNA synthesis so that DNA to cell mass is maintained at a constant ratio during cell division and DNA repair. A somewhat unusual feature of the RNR enzyme is that it catalyzes a reaction that proceeds via a free radical mechanism of action. The substrates for RNR are ADP, GDP, CDP, and UDP. dTDP (deoxythymidine diphosphate) is synthesized by another enzyme (thymidylate kinase) from dTMP (deoxythymidine monophosphate). The iron-dependent enzyme, ribonucleotide reductase (RNR), is essential for DNA synthesis. RNRs provide an essential link between the RNA and DNA world. That brings us to the classic chicken and egg, catch22 situation. RNR enzymes are required to make DNA. DNA is however required to make RNR enzymes. What came first ?? We can conclude with high certainty that this enzyme buries any RNA world fantasies, and any possibility of transition from RNA to DNA world scenarios. 21
Thymine
DNA can only be replicated in the presence of specific enzymes (described below ) which can only be manufactured by the already existing DNA. Each is absolutely essential for the other, and both must be present for the DNA to multiply. Therefore, DNA has to have been in existence at the beginning for life to be controlled by DNA. Scott M. Huse, "The Collapse of Evolution", Baker Book House: Grand Rapids (Michigan), 1983 p:93-94
That's interesting, as we find two distinct enzymes with two different sequences and structures synthesizing the same reaction, thus being a example of convergence right in the beginning. How remote was the chance for this to happen by natural means, considering, that convergence does not favor naturalistic explanations?
As Stephen J.Gould wrote: “…No finale can be specified at the start, none would ever occur a second time in the same way, because any pathway proceeds through thousands of improbable stages. Alter any early event, ever so slightly, and without apparent importance at the time, and evolution cascades into a radically different channel.1
Stephen J. Gould, Wonderful Life: The Burgess Shale and the Nature of History (New York, NY: W.W. Norton & Company, 1989), 51.
By virtue of their function and phyletic distribution, Thys are ancient enzymes, implying 1) the likely participation of one or both enzymes during the transition from an RNA world to a DNA world (based on protein catalysts: Joyce 2002) and 2) the probable presence of a gene encoding Thy in the genome of the common ancestors of eukaryotes, bacteria, and archaea . Thus, tracing back the pathway of genes encoding ThyA and ThyX may shed light on the actively debated wider issue regarding the origins of viral and cellular DNA
This brings us to the same problem as with Ribonucleotide Reductase enzymes (RNR), which is the classic chicken and egg, catch22 situation. ThyA and ThyX enzymes are required to make DNA. DNA is however required to make these enzymes. What came first ?? We can conclude with high certainty that this enzyme buries any RNA world fantasies, and any possibility of transition from RNA to DNA world scenarios, since both had to come into existence at the same time.
Which came first, proteins or protein synthesis?
Both the transcription and translation systems depend upon numerous proteins, many of which are jointly necessary for protein synthesis to occur at all. Yet all of these proteins are made by this very process. Proteins involved in transcription such as RNA polymerases, for example, are built from instructions carried on an RNA transcript. Translation of the RNA transcript depends upon other specialized enzymes such as synthetases, yet the information to build these enzymes is translated during the translation process that synthetases themselves facilitate. Biochemist David Goodsell describes the problem, "The key molecular process that makes modern life possible is protein synthesis, since proteins are used in nearly every aspect of living. The synthesis of proteins requires a tightly integrated sequence of reactions, most of which are themselves performed by proteins." Or as Jacques Monod noted in 1971: "The code is meaningless unless translated. The modern cell's translating machinery consists of at least fifty macromolecular components which are themselves coded in DNA: the code cannot be translated otherwise than by products of translation." (Scientists now know that translation actually requires more than a hundred proteins.) The integrated complexity of the cell's information-processing system has prompted some profound reflection. As Lewontin asks, "What makes the proteins that are necessary to make the protein?" As David Goodsell puts it, this "is one of the unanswered riddles of biochemistry: which came first, proteins or protein synthesis? If proteins are needed to make proteins, how did the whole thing get started?" The end result of protein synthesis is required before it can begin.
The Interdependency of Lipid Membranes and Membrane Proteins
The cell membrane contains various types of proteins, including ion channel proteins, proton pumps, G proteins, and enzymes. These membrane proteins function cooperatively to allow ions to penetrate the lipid bilayer. The interdependency of lipid membranes and membrane proteins suggests that lipid bilayers and membrane proteins co-evolved together with membrane bioenergetics. The nonsense of this assertion is evident. How could the membrane proteins co-evolve, if they had to be manufactured in the machinery, protected by the cell membrane? The cell membrane contains various types of proteins, including ion channel proteins, proton pumps, G proteins, and enzymes. These membrane proteins function cooperatively to allow ions to penetrate the lipid bilayer. The ER and Golgi apparatus together constitute the endomembrane compartment in the cytoplasm of eukaryotic cells. The endomembrane compartment is a major site of lipid synthesis, and the ER is where not only lipids are synthesized, but membrane-bound proteins and secretory proteins are also made.
So in order to make cell membranes, the Endoplasmic Reticulum is required. But also the Golgi Apparatus, the peroxisome, and the mitochondria. But these only function, if protected and encapsulated in the cell membrane. What came first, the cell membrane, or the endoplasmic reticulum? This is one of many other catch22 situations in the cell, which indicate that the cell could not emerge in a stepwise gradual manner, as proponents of natural mechanisms want to make us believe.
Not only is the cell membrane intricate and complex (and certainly not random), but it has tuning parameters such as the degree to which the phospholipid tails are saturated. It is another example of a sophisticated biological design about which evolutionists can only speculate. Random mutations must have luckily assembled molecular mechanisms which sense environmental challenges and respond to them by altering the phospholipid population in the membrane in just the right way. Such designs are tremendously helpful so of course, they would have been preserved by natural selection. It is yet another example of how silly evolutionary theory is in light of scientific facts. 16
The DNA - Enzyme System is Irreducibly Complex
An often undiscussed aspect of complexity is how the tRNA get assigned to the right amino acids. For the DNA language to be translated properly, each tRNA codon must be attached to the correct amino acid. If this crucial step in DNA replication is not functional, then the language of DNA breaks down. Special enzymes called aminoacyl - tRNA synthetases (aaRSs) ensure that the proper amino acid is attached to a tRNA with the correct codon through a chemical reaction called "aminoacylation." Accurate translation requires not only that each tRNA be assigned the correct amino acid, but also that it not be aminoacylated by any of the aaRS molecules for the other 19 amino acids. One biochemistry textbook notes that because all aaRSs catalyze similar reactions upon various similar tRNA molecules, it was thought they "evolved from a common ancestor and should therefore be structurally related." (Voet and Voet pg. 971-975) However, this was not the case as the, "aaRSs form a diverse group of [over 100] enzymes … and there is little sequence similarity among synthetases specific for different amino acids." (Voet and Voet pg. 971-975) Amazingly, these aaRSs themselves are coded for by the DNA: this forms the essence of a chicken-egg problem. The enzymes themselves build help perform the very task which constructs them! 9
Which came first, glycolysis to make energy or energy from glycolysis needed to make enzymes? Without the enzymes, glycolysis could not occur to produce ATP. But without the ATP those enzymes could not be manufactured. 10
mRNA is needed to make the nuclear pore complex. But without the nuclear pore complex, mRNA cannot be prepared for translation in the Ribosome. Thats a catch22 situation....1
Chicken and Egg
Yarus's model also raises a significant chicken-and-egg paradox. Meyer and Nelson explain: Because those biosynthetic pathways involve many enzymes, extant cells would require a pre-existing translation system in order to make them. Since attempts to explain the origin of the genetic code also attempt to explain the origin of the translation system (indeed, there can be no translation without a code), Yarus et al.'s findings raise an acute chicken and egg problem. Which came first, the aptamer-amino acid affinities that Yarus et al. propose as the basis of the code and translation system, or the translation system that would have been necessary to produce those amino acids (and, thus aptamer-amino acid affinities) in the first place? 2
A new chicken-and-egg paradox relating to the origin of life 12
Cells could not have evolved without viruses, as they need reverse transcriptase (which is found only in viruses) in order to move from RNA to DNA. In other words, instead of helping to solve the problem of the origin of life on Earth, recent research has only served to highlight one of its central paradoxes. And yet the science media reports the latest discoveries as if the solution is just around the corner. Don’t you find that just a little strange? “In order to move from RNA to DNA, you need an enzyme called reverse transcriptase,” Dolja said. “It’s only found in viruses like HIV, not in cells. So how could cells begin to use DNA without the help of a virus?” 13
The creation of double-stranded DNA occurs in the cytosol as a series of these steps:
A specific cellular tRNA acts as a primer and hybridizes to a complementary part of the virus RNA genome called the primer binding site or PBS
Complementary DNA then binds to the U5 (non-coding region) and R region (a direct repeat found at both ends of the RNA molecule) of the viral RNA
A domain on the reverse transcriptase enzyme called RNAse H degrades the 5’ end of the RNA which removes the U5 and R region
The primer then ‘jumps’ to the 3’ end of the viral genome and the newly synthesized DNA strands hybridize to the complementary R region on the RNA
The first strand of complementary DNA (cDNA) is extended and the majority of viral RNA is degraded by RNAse H
Once the strand is completed, second strand synthesis is initiated from the viral RNA
There is then another ‘jump’ where the PBS from the second strand hybridizes with the complementary PBS on the first strand
Both strands are extended further and can be incorporated into the host's genome by the enzyme integrase
Proteins are required to make proteins 14
The threat of autodestruction stems from the circular nature of protein synthesis. Proteins constitute many components of the cell's protein manufacturing machinery. In other words, the cell uses proteins to make proteins. So, if the protein manufacturing machinery were assembled with defective parts, the cell would fail to accurately manufacture proteins. Such a manufacturing failure would cause protein production systems to become increasingly error-prone with each successive round of protein synthesis. Protein manufacturing systems made up of defective components would be more likely to produce defective proteins. This chain reaction would cascade out of control and quite quickly lead to the cells self-destruction. Effective quality assurance procedures must be in place for protein production or life would not be possible.
Creation of double-stranded DNA also involves strand transfer, in which there is a translocation of short DNA product from initial RNA-dependent DNA synthesis to acceptor template regions at the other end of the genome, which are later reached and processed by the reverse transcriptase for its DNA-dependent DNA activity
The process of DNA replication depends on many separate protein catalysts to unwind, stabilize, copy, edit, and rewind the original DNA message. In prokaryotic cells, DNA replication involves more than thirty specialized proteins to perform tasks necessary for building and accurately copying the genetic molecule. These specialized proteins include DNA polymerases, primases, helicases, topoisomerases, DNA-binding proteins, DNA ligases, and editing enzymes.38 DNA needs these proteins to copy the genetic information contained in DNA. But the proteins that copy the genetic information in DNA are themselves built from that information. This again poses what is, at the very least, a curiosity: the production of proteins requires DNA, but the production of DNA requires proteins.
“The ‘Catch-22’ of the origin of life is this: DNA can replicate, but it needs enzymes in order to catalyze the process. Proteins can catalyse DNA formation, but they need DNA to specify the correct sequence of amino acids.” Indeed, the origin of the genetic code is a vicious circle: protein machines are needed to read the DNA, but these protein machines are themselves encoded on the DNA. Furthermore, they use energy, which requires ATP, made by the nano-motor ATP synthase. Yet this is encoded on the DNA, decoded by machines needing ATP. The proteins are the machinery, and the DNA is the reproductive material, yet both are needed at the same time for the cell to function at all. And of course, this would be useless without any information to reproduce.
Nar1 is both a target and a component of the cellular Fe/S protein biogenesis machinery creating an interesting “chicken and egg” situation for its maturation process (Balk et al., 2004). 5
How on earth could proofreading enzymes emerge, especially with this degree of fidelity, when they depend on the very information that they are designed to protect? Think about it. This is a catch-22 for Darwinists. 6
Koonin, the logic of chance, page 231:
So an inevitable (even if perhaps counterintuitive) conclusion from the comparative analysis of ancient paralogous relationship between protein components of the translation system is that, with the interesting exception of the core ribosomal proteins, all proteins that play essential roles in modern translation are products of a long and complex evolution of diverse protein domains. Here comes the Catch-22: For all this protein evolution to occur, an accurate and efficient translation system is required. This primordial translation system might not need to be quite as good as the modern version, but it seems a safe bet that is must have been within an order of magnitude from the modern one in terms of fidelity and translation rates to make protein evolution possible. However, from all we know about the modern translation system, this level of precision is unimaginable without a complex, dedicated protein apparatus. Thus, the translation system presents us with the “Darwin-Eigen paradox” that is inherent to all thinking on the emergence of complex biological entities: For a modern-type, efficient, and accurate
translation system to function, many diverse proteins are required, but for those proteins to evolve, a translation system almost as good as the modern one would be necessary.
RNA synthesis requires RNA repair enzymes
A cell has a great investment in its RNAs – they are working copies of its genomic information. The study of mRNA biogenesis in the last few years has revealed an elaborate surveillance mechanism involving factors such as the UPF proteins that culls aberrantly spliced mRNAs and mRNAs with premature termination codons. There might be a hint that such RNA quality control mechanisms go awry in cancers, just as DNA quality control mechanisms do, where aberrantly spliced transcripts accumulate in a tumor. Now that the gates are open, we may have a flood of studies on the RNome [the RNA genome] stability and cancer.
This aggravates the chicken-and-egg problem for proponents of natural mechanisms. In the “RNA World” hypothesis for the origin of life, RNA performed both the information storage and enzymatic functions before these roles were outsourced to DNA and proteins. But how could RNA repair itself? If RNA needs to be protected from damage, the protein repair system would have needed to be there from the beginning. Proponents of natural mechanisms might surmise that different primitive RNAs worked side by side to repair each other, but that strains credibility for a hypothesis is already far-fetched. In typical evolutionary lingo, Begley and Samson blow smoke about what nature produced (emphasis added): “It seems that, for each human protein, parameters have evolved to distinguish between RNA and DNA,” they speculate, and in another place, “It might be that the RNA-demethylation activity of AlkB-like proteins evolved to regulate biological RNA methylation, and that the repair of aberrant, chemical methylation is fortuitous.” Ask them how the cell evolved these things, and you’ll probably get a quizzical look, as if “Why are you asking such a dumb question? I don’t know. It just had to. We’re here, aren’t we?” 7
Recombination Vital to Genome Stability 8
The latest issue of the Proceedings of the National Academy of Sciences (July 17) contains a symposium on gene replication and recombination, among other papers on DNA. Among the interesting papers:
(1) A theory on how genomes can contain vast stretches of non-coding DNA, apparently inactive retrotransposons that were inserted by recombination, polyploidy or lateral transfer. These inactive stretches, while harmless, can greatly expand the genome while keeping the number of actual genes relatively constant.
(2) A description of how recombination is an essential method for repair of DNA breaks, stating that “DNA synthesis is an accurate and very processive phenomenon; nevertheless, replication fork progression on chromosomes can be impeded by DNA lesions, DNA secondary structures, or DNA-bound proteins. Elements interfering with the progression of replication forks have been reported to induce rearrangements and/or render homologous recombination essential for viability, in all organisms from bacteria to human.”
(3) Another paper describes how specialized proteins called topoisomerases help prevent the strain of uncoiling DNA from breaking, but when they fail, recombination can help restart the replication process.
(4) A paper describes how recombination works to repair breaks in a replicating chromosome.
(5) Some Japanese scientists describe how a gene codes for a motor protein that is essential for genome stability.
(6) The cover story describes the various repair mechanisms, stating, “Maintenance of genomic integrity and stable transmission of genetic information depend on a number of DNA repair processes. Failure to faithfully perform these processes can result in genetic alterations and subsequent development of cancer and other genetic diseases.” Describing one such mechanism named Rad52, the authors state, “The key role played by Rad52 in this pathway has been attributed to its ability to seek out and mediate annealing of homologous DNA strands . . . . our data indicate that each Rad52 focus [i.e. active site] represents a center of recombinational repair capable of processing multiple DNA lesions.”
These are just samples of the exciting findings being made about DNA replication. These and other papers show that it is a fail-safe system with many sophisticated backup and repair mechanisms. While there is still much to learn, and many mysteries to explain, DNA’s ability to replicate is truly a marvel of engineering. Think about the classic chicken-and-egg conundrum for evolution illustrated by (5) above: a gene codes for a protein that is essential for the gene to exist. Browse through the abstracts of these papers just to get a feel for the amazingly complex world of cellular processes going on in your body right now, without your conscious thought or control.
You need energy to make energy
once Earth had pyrophosphite, it had an energetic molecule that, while not as useful as ATP, was at least somewhat similar. “The team found that a compound known as pyrophosphite may have been an important energy source for primitive lifeforms.” Did he have any evidence for this? No; it’s just a requirement. “It’s a chicken and egg question,” he said. “Scientists are in disagreement over what came first – replication, or metabolism. But there is a third part to the equation – and that is energy.” So while scientists are disagreeing about two things, why not add a third? That’s progress: “You need enzymes to make ATP and you need ATP to make enzymes,” explained Dr Kee, as if adding questions qualifies as explaining something: “The question is: where did energy come from before either of these two things existed?” We may not know the answers, but at least our ignorance is getting more sophisticated thanks to OOL research.17
The hardware and software of the cell, evidence of design 18
Paul Davies: the fifth miracle page 62
Due to the organizational structure of systems capable of processing algorithmic (instructional) information, it is not at all clear that a monomolecular system – where a single polymer plays the role of catalyst and informational carrier – is even logically consistent with the organization of information flow in living systems, because there is no possibility of separating information storage from information processing (that being such a distinctive feature of modern life). As such, digital-first systems (as currently posed) represent a rather trivial form of information processing that fails to capture the logical structure of life as we know it. 1
We need to explain the origin of both the hardware and software aspects of life, or the job is only half finished. Explaining the chemical substrate of life and claiming it as a solution to life’s origin is like pointing to silicon and copper as an explanation for the goings-on inside a computer. It is this transition where one should expect to see a chemical system literally take-on “a life of its own”, characterized by informational dynamics which become decoupled from the dictates of local chemistry alone (while of course remaining fully consistent with those dictates). Thus the famed chicken-or-egg problem (a solely hardware issue) is not the true sticking point. Rather, the puzzle lies with something fundamentally different, a problem of causal organization having to do with the separation of informational and mechanical aspects into parallel causal narratives. The real challenge of life’s origin is thus to explain how instructional information control systems emerge naturally and spontaneously from mere molecular dynamics.
Software and hardware are irreducibly complex and interdependent. There is no reason for information processing machinery to exist without the software, and vice versa.
Systems of interconnected software and hardware are irreducibly complex.
All cellular functions are irreducibly complex
Paul Davies, the fifth miracle page 53:
Pluck the DNA from a living cell and it would be stranded, unable to carry out its familiar role. Only within the context of a highly specific molecular milieu will a given molecule play its role in life. To function properly, DNA must be part of a large team, with each molecule executing its assigned task alongside the others in a cooperative manner. Acknowledging the interdependability of the component molecules within a living organism immediately presents us with a stark philosophical puzzle. If everything needs everything else, how did the community of molecules ever arise in the first place? Since most large molecules needed for life are produced only by living organisms, and are not found outside the cell, how did they come to exist originally, without the help of a meddling scientist? Could we seriously expect a Miller-Urey type of soup to make them all at once, given the hit-and-miss nature of its chemistry? 19
Being part of a large team, cooperative manner, inter dependability, everything needs everything else, are just other words for irreducibility and interdependence.
For a nonliving system, questions about irreducible complexity are even more challenging for a totally natural non-design scenario, because natural selection — which is the main mechanism of Darwinian evolution — cannot exist until a system can reproduce. For the origin of life, we can think about the minimal complexity that would be required for reproduction and other basic life functions. Most scientists think this would require hundreds of biomolecular parts, not just the five parts in a simple mousetrap or in my imaginary LMNOP system. And current science has no plausible theories to explain how the minimal complexity required for life (and the beginning of biological natural selection) could have been produced by natural processes before the beginning of biological natural selection.
While the heart serves as a “pump” to deliver blood throughout the body, it also requires oxygenated blood in order to remain healthy. Or consider that the production of red blood cells, one of the major constituents within the vascular system, is regulated by erythropoietin—a hormone produced in the kidney. Yet that kidney requires red blood cells to deliver oxygenated blood. So which evolved first? 22
The Interdependency of Lipid Membranes and Membrane Proteins
Even in the simplest cells, the membrane is a biological device of a staggering complexity that carries diverse protein complexes mediating energy-dependent – and tightly regulated - import and export of metabolites and polymers
Remarkably, even the author of the book: Agents Under Fire: Materialism and the Rationality of Science, pgs. 104-105 (Rowman & Littlefield, 2004). HT: ENV. asks the readers: Hence a chicken and egg paradox: a lipid membrane would be useless without membrane proteins but how could membrane proteins have evolved in the absence of functional membranes?
Biosynthesis of cell membranes: It takes membranes to make membranes
Cell walls indeed provide essential structural support and external interactions in modern organisms (Albers & Meyer, 2011), Despite the stunning diversity that exists among prokaryotic cell envelopes, the synthesis of many of their main components:
1.N-or O-glycosylated S-layer proteins
2.peptidoglycan,
3.O-antigen LPS
4.teichoic acids
5.exopolysaccharides
relies on comparable glycosylation pathways These pathways are all located in the cell membranes, are mediated by similar lipid carriers and have the same orientation across the membrane.
That raises the question: If in extant cells these pathways are located in the cell membranes, where were to location prior membranes existed ? Would it not be correct to say : It takes membranes to make membranes ?
1) https://reasonandscience.catsboard.com/t2117-nuclear-pores#3762
2) http://www.evolutionnews.org/2011/08/direct_rna_templating_a_failed050121.html
3) http://www.fis.puc.cl/~jalfaro/astrobiologia/Astrobiologiavasquez.pdf
4) Meyer, signature of the cell, page 111
5) https://reasonandscience.catsboard.com/t2285-iron-sulfur-clusters-basic-building-blocks-for-life#4646
6) https://reasonandscience.catsboard.com/t2043-dna-error-checking-and-repair#4669
7) https://reasonandscience.catsboard.com/t2043-dna-and-rna-error-checking-and-repair-amazing-evidence-of-design#4671
8 https://reasonandscience.catsboard.com/t1849p30-dna-replication-of-prokaryotes#4672
9) http://www.ideacenter.org/contentmgr/showdetails.php/id/845
10) https://reasonandscience.catsboard.com/t1796-glycolysis?highlight=glycolysis
11) http://www.scientificamerican.com/article/a-simpler-origin-for-life/
12) http://www.uncommondescent.com/intelligent-design/do-viruses-help-explain-the-origin-of-life/
13) https://en.wikipedia.org/wiki/Reverse_transcriptase#In_eukaryotes
14) Fazale Rana, Cell's design, page 186
16) https://reasonandscience.catsboard.com/t1331-the-cell-membrane-irreducible-complexity
17) http://creationsafaris.com/crev201005.htm
18) https://reasonandscience.catsboard.com/t2221-the-hardware-and-software-of-the-cell-evidence-of-design?highlight=software
19) https://reasonandscience.catsboard.com/t2179-the-cell-is-a-interdependent-irreducible-complex-system
20) https://reasonandscience.catsboard.com/t2285-iron-sulfur-clusters-basic-building-blocks-for-life
21) https://reasonandscience.catsboard.com/t2028-origin-of-the-dna-double-helix#3426
22) http://www.apologeticspress.org/ApPubPage.aspx?pub=1&issue=571&article=450
23) https://reasonandscience.catsboard.com/t2371-how-cellular-enzymatic-and-metabolic-networks-point-to-design#5139
24) https://www.quantamagazine.org/lifes-first-molecule-was-protein-not-rna-new-model-suggests-20171102/
https://reasonandscience.catsboard.com/t2059-catch22-chicken-and-egg-problems-in-biology-and-biochemistry

Life’s First Molecule Was Protein, Not RNA, New Model Suggests
For scientists studying the origin of life, one of the greatest chicken-or-the-egg questions is: Which came first — proteins or nucleic acids like DNA and RNA? 24
Information is required to extract energy. But extract energy is required to store information. What came first?
The Catch22 riddle in 1965
https://sci-hub.ren/https://www.nature.com/articles/205328a0
what changed, is that rather than closing the gaps, they have become wider and wider, and naturalistic proposals more and more unrealistic!
In Joseph Heller’s classic novel about World War II,Catch-22, an aviator could be excused from combat duty for being crazy. But a rule specified that he first had to request an excuse, and anyone who requested an excuse from combat duty was obviously not crazy, so such requests were invariably denied. The rule that made it impossible to be excused from combat duty was called “Catch-22.”

Oxygen and life have a catch 22 relationship
Catch 22 is a situation in which an action has consequences which make impossible to pursue that action. Oxygen is very harmful to life. At the same time oxygen is needed to provide the ozone layer which protects life from ultraviolet radiation (UVR) coming from the sun. If Cyanobacteria came before oxygen, because it is the cause of oxygen, then Cyanobacteria would have had to develop several forms of protection to mitigate the damage from UVR: avoidance, scavenging, screening, repair, and programmed cell death. However, UVR damage is immediate and the time needed to “evolve” protection against it via natural selection, incredibly slow. So, UVR damage would occur before any such defense mechanisms could evolve. 3
Iron-sulfur clusters
Sulfur is an essential element, being a constituent of many proteins and cofactors. Iron-sulfur (FeS) centers are essential protein cofactors in all forms of life. Various biosynthetic pathways were found to be tightly interconnected through complex crosstalk mechanisms that crucially depend on the bio-availability of the metal ions iron, molybdenum, tungsten, nickel, copper, and zinc. Proteins requiring Fe/S clusters in their active site have been localized in mitochondria, cytosol, and nucleus where they are involved in rather diverse functions such as the TCA cycle, amino acid biosynthesis, bacterial and mitochondrial respiration, co-factor biosynthesis, ribosome assembly, regulation of protein translation, DNA replication and DNA repair. Hence the process of iron-sulfur biosynthesis is essential to almost all forms of life. The prevalence of these proteins on the metabolic pathways of most organisms leads some scientists to theorize that iron-sulfur compounds had a significant role in the origin of life in the iron-sulfur world theory. The iron-sulfur world hypothesis is a set of proposals for the origin of life and the early evolution of life advanced in a series of articles between 1988 and 1992 by Günter Wächtershäuser. FeS cluster assembly is a complex process involving the mobilization of Fe and S atoms from storage sources, their assembly into [Fe-S] form, their transport to specific cellular locations, and their transfer to recipient apoproteins. Nar1 is an essential component of a cytosolic Fe/S protein assembly machinery. Required for maturation of extramitochondrial Fe/S proteins. 12 Thus, Nar1 is both a target and a component of the cellular Fe/S protein biogenesis machinery creating an interesting “chicken and egg” situation for its maturation process Conserved Iron–Sulfur (Fe–S) clusters are found in a growing family of metalloproteins that are implicated in prokaryotic and eukaryotic DNA replication and repair. 20
Therefore, they had to exist prior to life beginning, since DNA replication enzymes and proteins depend on them. They require however also complex proteins and enzymes to be synthesized. That's a classical chicken/egg problem.
Ribonucleotide reductase
This is one of the most essential enzymes of life. Ribonucleotide reduction is the only pathway for de novo synthesis of deoxyribonucleotides in extant organisms. This chemically demanding reaction is catalyzed by ribonucleotide reductase (RNR). The mechanism has been deemed unlikely to be catalyzed by a ribozyme, creating an enigma regarding how the building blocks for DNA were synthesized at the transition from RNA to DNA-encoded genomes. Biosynthesis DNA is made from RNA. The deoxynucleotides are made from nucleotides with ribonucleotide reductases (RNRs), producing uracil-DNA or u-DNA. The uracil is then converted to thymine by adding a methyl group, making thymine-DNA or t-DNA, the kind that is actually used. The reaction catalyzed by RNR is strictly conserved in all living organisms. Furthermore, RNR plays a critical role in regulating the total rate of DNA synthesis so that DNA to cell mass is maintained at a constant ratio during cell division and DNA repair. A somewhat unusual feature of the RNR enzyme is that it catalyzes a reaction that proceeds via a free radical mechanism of action. The substrates for RNR are ADP, GDP, CDP, and UDP. dTDP (deoxythymidine diphosphate) is synthesized by another enzyme (thymidylate kinase) from dTMP (deoxythymidine monophosphate). The iron-dependent enzyme, ribonucleotide reductase (RNR), is essential for DNA synthesis. RNRs provide an essential link between the RNA and DNA world. That brings us to the classic chicken and egg, catch22 situation. RNR enzymes are required to make DNA. DNA is however required to make RNR enzymes. What came first ?? We can conclude with high certainty that this enzyme buries any RNA world fantasies, and any possibility of transition from RNA to DNA world scenarios. 21
Thymine
DNA can only be replicated in the presence of specific enzymes (described below ) which can only be manufactured by the already existing DNA. Each is absolutely essential for the other, and both must be present for the DNA to multiply. Therefore, DNA has to have been in existence at the beginning for life to be controlled by DNA. Scott M. Huse, "The Collapse of Evolution", Baker Book House: Grand Rapids (Michigan), 1983 p:93-94
Thymidylate synthases (Thy) are key enzymes in the synthesis of deoxythymidylate, 1 of the 4 building blocks of DNA. As such, they are essential for all DNA-based forms of life and therefore implicated in the hypothesized transition from RNA genomes to DNA genomes. Two unrelated Thy enzymes, ThyA and ThyX, are known to catalyze the same biochemical reaction.
Thymidylate synthase (Thy) is a fundamental enzyme in DNA synthesis because it catalyzes the formation of deoxythymidine 5′-monophosphate (dTMP) from deoxyuridine 5′-monophosphate (dUMP). For decades, only one family of thymidylate synthase enzymes was known, and its presence was considered necessary to maintain all DNA-based forms of life. Then, a gene encoding an alternative enzyme was discovered and characterized, and the novel enzyme was named ThyX, whereas the other enzyme was renamedThyA. Even though both reactions accomplish the same key step, the reaction mechanisms or steps, catalyzed by the FDTS and TS enzymes are structurally different. The 2 enzymes, ThyA and ThyX, were found to have distinctly different sequences and structures, thus alluding to independent origins. 21
Thymidylate synthase (Thy) is a fundamental enzyme in DNA synthesis because it catalyzes the formation of deoxythymidine 5′-monophosphate (dTMP) from deoxyuridine 5′-monophosphate (dUMP). For decades, only one family of thymidylate synthase enzymes was known, and its presence was considered necessary to maintain all DNA-based forms of life. Then, a gene encoding an alternative enzyme was discovered and characterized, and the novel enzyme was named ThyX, whereas the other enzyme was renamedThyA. Even though both reactions accomplish the same key step, the reaction mechanisms or steps, catalyzed by the FDTS and TS enzymes are structurally different. The 2 enzymes, ThyA and ThyX, were found to have distinctly different sequences and structures, thus alluding to independent origins. 21
That's interesting, as we find two distinct enzymes with two different sequences and structures synthesizing the same reaction, thus being a example of convergence right in the beginning. How remote was the chance for this to happen by natural means, considering, that convergence does not favor naturalistic explanations?
As Stephen J.Gould wrote: “…No finale can be specified at the start, none would ever occur a second time in the same way, because any pathway proceeds through thousands of improbable stages. Alter any early event, ever so slightly, and without apparent importance at the time, and evolution cascades into a radically different channel.1
Stephen J. Gould, Wonderful Life: The Burgess Shale and the Nature of History (New York, NY: W.W. Norton & Company, 1989), 51.
By virtue of their function and phyletic distribution, Thys are ancient enzymes, implying 1) the likely participation of one or both enzymes during the transition from an RNA world to a DNA world (based on protein catalysts: Joyce 2002) and 2) the probable presence of a gene encoding Thy in the genome of the common ancestors of eukaryotes, bacteria, and archaea . Thus, tracing back the pathway of genes encoding ThyA and ThyX may shed light on the actively debated wider issue regarding the origins of viral and cellular DNA
This brings us to the same problem as with Ribonucleotide Reductase enzymes (RNR), which is the classic chicken and egg, catch22 situation. ThyA and ThyX enzymes are required to make DNA. DNA is however required to make these enzymes. What came first ?? We can conclude with high certainty that this enzyme buries any RNA world fantasies, and any possibility of transition from RNA to DNA world scenarios, since both had to come into existence at the same time.
Which came first, proteins or protein synthesis?
Both the transcription and translation systems depend upon numerous proteins, many of which are jointly necessary for protein synthesis to occur at all. Yet all of these proteins are made by this very process. Proteins involved in transcription such as RNA polymerases, for example, are built from instructions carried on an RNA transcript. Translation of the RNA transcript depends upon other specialized enzymes such as synthetases, yet the information to build these enzymes is translated during the translation process that synthetases themselves facilitate. Biochemist David Goodsell describes the problem, "The key molecular process that makes modern life possible is protein synthesis, since proteins are used in nearly every aspect of living. The synthesis of proteins requires a tightly integrated sequence of reactions, most of which are themselves performed by proteins." Or as Jacques Monod noted in 1971: "The code is meaningless unless translated. The modern cell's translating machinery consists of at least fifty macromolecular components which are themselves coded in DNA: the code cannot be translated otherwise than by products of translation." (Scientists now know that translation actually requires more than a hundred proteins.) The integrated complexity of the cell's information-processing system has prompted some profound reflection. As Lewontin asks, "What makes the proteins that are necessary to make the protein?" As David Goodsell puts it, this "is one of the unanswered riddles of biochemistry: which came first, proteins or protein synthesis? If proteins are needed to make proteins, how did the whole thing get started?" The end result of protein synthesis is required before it can begin.
The Interdependency of Lipid Membranes and Membrane Proteins
The cell membrane contains various types of proteins, including ion channel proteins, proton pumps, G proteins, and enzymes. These membrane proteins function cooperatively to allow ions to penetrate the lipid bilayer. The interdependency of lipid membranes and membrane proteins suggests that lipid bilayers and membrane proteins co-evolved together with membrane bioenergetics. The nonsense of this assertion is evident. How could the membrane proteins co-evolve, if they had to be manufactured in the machinery, protected by the cell membrane? The cell membrane contains various types of proteins, including ion channel proteins, proton pumps, G proteins, and enzymes. These membrane proteins function cooperatively to allow ions to penetrate the lipid bilayer. The ER and Golgi apparatus together constitute the endomembrane compartment in the cytoplasm of eukaryotic cells. The endomembrane compartment is a major site of lipid synthesis, and the ER is where not only lipids are synthesized, but membrane-bound proteins and secretory proteins are also made.
So in order to make cell membranes, the Endoplasmic Reticulum is required. But also the Golgi Apparatus, the peroxisome, and the mitochondria. But these only function, if protected and encapsulated in the cell membrane. What came first, the cell membrane, or the endoplasmic reticulum? This is one of many other catch22 situations in the cell, which indicate that the cell could not emerge in a stepwise gradual manner, as proponents of natural mechanisms want to make us believe.
Not only is the cell membrane intricate and complex (and certainly not random), but it has tuning parameters such as the degree to which the phospholipid tails are saturated. It is another example of a sophisticated biological design about which evolutionists can only speculate. Random mutations must have luckily assembled molecular mechanisms which sense environmental challenges and respond to them by altering the phospholipid population in the membrane in just the right way. Such designs are tremendously helpful so of course, they would have been preserved by natural selection. It is yet another example of how silly evolutionary theory is in light of scientific facts. 16
The DNA - Enzyme System is Irreducibly Complex
An often undiscussed aspect of complexity is how the tRNA get assigned to the right amino acids. For the DNA language to be translated properly, each tRNA codon must be attached to the correct amino acid. If this crucial step in DNA replication is not functional, then the language of DNA breaks down. Special enzymes called aminoacyl - tRNA synthetases (aaRSs) ensure that the proper amino acid is attached to a tRNA with the correct codon through a chemical reaction called "aminoacylation." Accurate translation requires not only that each tRNA be assigned the correct amino acid, but also that it not be aminoacylated by any of the aaRS molecules for the other 19 amino acids. One biochemistry textbook notes that because all aaRSs catalyze similar reactions upon various similar tRNA molecules, it was thought they "evolved from a common ancestor and should therefore be structurally related." (Voet and Voet pg. 971-975) However, this was not the case as the, "aaRSs form a diverse group of [over 100] enzymes … and there is little sequence similarity among synthetases specific for different amino acids." (Voet and Voet pg. 971-975) Amazingly, these aaRSs themselves are coded for by the DNA: this forms the essence of a chicken-egg problem. The enzymes themselves build help perform the very task which constructs them! 9
Which came first, glycolysis to make energy or energy from glycolysis needed to make enzymes? Without the enzymes, glycolysis could not occur to produce ATP. But without the ATP those enzymes could not be manufactured. 10
A Simpler Origin for Life
DNA replication cannot proceed without the assistance of a number of proteins--members of a family of large molecules that are chemically very different from DNA. Proteins, like DNA, are constructed by linking subunits, amino acids in this case, together to form a long chain. Cells employ twenty of these building blocks in the proteins that they make, affording a variety of products capable of performing many different tasks--proteins are the handymen of the living cell. Their most famous subclass, the enzymes, act as expeditors, speeding up chemical processes that would otherwise take place too slowly to be of use to life. The above account brings to mind the old riddle: Which came first, the chicken or the egg? DNA holds the recipe for protein construction. Yet that information cannot be retrieved or copied without the assistance of proteins. Which large molecule, then, appeared first in getting life started--proteins (the chicken) or DNA (the egg)? 11mRNA is needed to make the nuclear pore complex. But without the nuclear pore complex, mRNA cannot be prepared for translation in the Ribosome. Thats a catch22 situation....1
Chicken and Egg
Yarus's model also raises a significant chicken-and-egg paradox. Meyer and Nelson explain: Because those biosynthetic pathways involve many enzymes, extant cells would require a pre-existing translation system in order to make them. Since attempts to explain the origin of the genetic code also attempt to explain the origin of the translation system (indeed, there can be no translation without a code), Yarus et al.'s findings raise an acute chicken and egg problem. Which came first, the aptamer-amino acid affinities that Yarus et al. propose as the basis of the code and translation system, or the translation system that would have been necessary to produce those amino acids (and, thus aptamer-amino acid affinities) in the first place? 2
A new chicken-and-egg paradox relating to the origin of life 12
Cells could not have evolved without viruses, as they need reverse transcriptase (which is found only in viruses) in order to move from RNA to DNA. In other words, instead of helping to solve the problem of the origin of life on Earth, recent research has only served to highlight one of its central paradoxes. And yet the science media reports the latest discoveries as if the solution is just around the corner. Don’t you find that just a little strange? “In order to move from RNA to DNA, you need an enzyme called reverse transcriptase,” Dolja said. “It’s only found in viruses like HIV, not in cells. So how could cells begin to use DNA without the help of a virus?” 13
The creation of double-stranded DNA occurs in the cytosol as a series of these steps:
A specific cellular tRNA acts as a primer and hybridizes to a complementary part of the virus RNA genome called the primer binding site or PBS
Complementary DNA then binds to the U5 (non-coding region) and R region (a direct repeat found at both ends of the RNA molecule) of the viral RNA
A domain on the reverse transcriptase enzyme called RNAse H degrades the 5’ end of the RNA which removes the U5 and R region
The primer then ‘jumps’ to the 3’ end of the viral genome and the newly synthesized DNA strands hybridize to the complementary R region on the RNA
The first strand of complementary DNA (cDNA) is extended and the majority of viral RNA is degraded by RNAse H
Once the strand is completed, second strand synthesis is initiated from the viral RNA
There is then another ‘jump’ where the PBS from the second strand hybridizes with the complementary PBS on the first strand
Both strands are extended further and can be incorporated into the host's genome by the enzyme integrase
Proteins are required to make proteins 14
The threat of autodestruction stems from the circular nature of protein synthesis. Proteins constitute many components of the cell's protein manufacturing machinery. In other words, the cell uses proteins to make proteins. So, if the protein manufacturing machinery were assembled with defective parts, the cell would fail to accurately manufacture proteins. Such a manufacturing failure would cause protein production systems to become increasingly error-prone with each successive round of protein synthesis. Protein manufacturing systems made up of defective components would be more likely to produce defective proteins. This chain reaction would cascade out of control and quite quickly lead to the cells self-destruction. Effective quality assurance procedures must be in place for protein production or life would not be possible.
Creation of double-stranded DNA also involves strand transfer, in which there is a translocation of short DNA product from initial RNA-dependent DNA synthesis to acceptor template regions at the other end of the genome, which are later reached and processed by the reverse transcriptase for its DNA-dependent DNA activity
The process of DNA replication depends on many separate protein catalysts to unwind, stabilize, copy, edit, and rewind the original DNA message. In prokaryotic cells, DNA replication involves more than thirty specialized proteins to perform tasks necessary for building and accurately copying the genetic molecule. These specialized proteins include DNA polymerases, primases, helicases, topoisomerases, DNA-binding proteins, DNA ligases, and editing enzymes.38 DNA needs these proteins to copy the genetic information contained in DNA. But the proteins that copy the genetic information in DNA are themselves built from that information. This again poses what is, at the very least, a curiosity: the production of proteins requires DNA, but the production of DNA requires proteins.
“The ‘Catch-22’ of the origin of life is this: DNA can replicate, but it needs enzymes in order to catalyze the process. Proteins can catalyse DNA formation, but they need DNA to specify the correct sequence of amino acids.” Indeed, the origin of the genetic code is a vicious circle: protein machines are needed to read the DNA, but these protein machines are themselves encoded on the DNA. Furthermore, they use energy, which requires ATP, made by the nano-motor ATP synthase. Yet this is encoded on the DNA, decoded by machines needing ATP. The proteins are the machinery, and the DNA is the reproductive material, yet both are needed at the same time for the cell to function at all. And of course, this would be useless without any information to reproduce.
Nar1 is both a target and a component of the cellular Fe/S protein biogenesis machinery creating an interesting “chicken and egg” situation for its maturation process (Balk et al., 2004). 5
How on earth could proofreading enzymes emerge, especially with this degree of fidelity, when they depend on the very information that they are designed to protect? Think about it. This is a catch-22 for Darwinists. 6
Koonin, the logic of chance, page 231:
So an inevitable (even if perhaps counterintuitive) conclusion from the comparative analysis of ancient paralogous relationship between protein components of the translation system is that, with the interesting exception of the core ribosomal proteins, all proteins that play essential roles in modern translation are products of a long and complex evolution of diverse protein domains. Here comes the Catch-22: For all this protein evolution to occur, an accurate and efficient translation system is required. This primordial translation system might not need to be quite as good as the modern version, but it seems a safe bet that is must have been within an order of magnitude from the modern one in terms of fidelity and translation rates to make protein evolution possible. However, from all we know about the modern translation system, this level of precision is unimaginable without a complex, dedicated protein apparatus. Thus, the translation system presents us with the “Darwin-Eigen paradox” that is inherent to all thinking on the emergence of complex biological entities: For a modern-type, efficient, and accurate
translation system to function, many diverse proteins are required, but for those proteins to evolve, a translation system almost as good as the modern one would be necessary.
RNA synthesis requires RNA repair enzymes
A cell has a great investment in its RNAs – they are working copies of its genomic information. The study of mRNA biogenesis in the last few years has revealed an elaborate surveillance mechanism involving factors such as the UPF proteins that culls aberrantly spliced mRNAs and mRNAs with premature termination codons. There might be a hint that such RNA quality control mechanisms go awry in cancers, just as DNA quality control mechanisms do, where aberrantly spliced transcripts accumulate in a tumor. Now that the gates are open, we may have a flood of studies on the RNome [the RNA genome] stability and cancer.
This aggravates the chicken-and-egg problem for proponents of natural mechanisms. In the “RNA World” hypothesis for the origin of life, RNA performed both the information storage and enzymatic functions before these roles were outsourced to DNA and proteins. But how could RNA repair itself? If RNA needs to be protected from damage, the protein repair system would have needed to be there from the beginning. Proponents of natural mechanisms might surmise that different primitive RNAs worked side by side to repair each other, but that strains credibility for a hypothesis is already far-fetched. In typical evolutionary lingo, Begley and Samson blow smoke about what nature produced (emphasis added): “It seems that, for each human protein, parameters have evolved to distinguish between RNA and DNA,” they speculate, and in another place, “It might be that the RNA-demethylation activity of AlkB-like proteins evolved to regulate biological RNA methylation, and that the repair of aberrant, chemical methylation is fortuitous.” Ask them how the cell evolved these things, and you’ll probably get a quizzical look, as if “Why are you asking such a dumb question? I don’t know. It just had to. We’re here, aren’t we?” 7
Recombination Vital to Genome Stability 8
The latest issue of the Proceedings of the National Academy of Sciences (July 17) contains a symposium on gene replication and recombination, among other papers on DNA. Among the interesting papers:
(1) A theory on how genomes can contain vast stretches of non-coding DNA, apparently inactive retrotransposons that were inserted by recombination, polyploidy or lateral transfer. These inactive stretches, while harmless, can greatly expand the genome while keeping the number of actual genes relatively constant.
(2) A description of how recombination is an essential method for repair of DNA breaks, stating that “DNA synthesis is an accurate and very processive phenomenon; nevertheless, replication fork progression on chromosomes can be impeded by DNA lesions, DNA secondary structures, or DNA-bound proteins. Elements interfering with the progression of replication forks have been reported to induce rearrangements and/or render homologous recombination essential for viability, in all organisms from bacteria to human.”
(3) Another paper describes how specialized proteins called topoisomerases help prevent the strain of uncoiling DNA from breaking, but when they fail, recombination can help restart the replication process.
(4) A paper describes how recombination works to repair breaks in a replicating chromosome.
(5) Some Japanese scientists describe how a gene codes for a motor protein that is essential for genome stability.
(6) The cover story describes the various repair mechanisms, stating, “Maintenance of genomic integrity and stable transmission of genetic information depend on a number of DNA repair processes. Failure to faithfully perform these processes can result in genetic alterations and subsequent development of cancer and other genetic diseases.” Describing one such mechanism named Rad52, the authors state, “The key role played by Rad52 in this pathway has been attributed to its ability to seek out and mediate annealing of homologous DNA strands . . . . our data indicate that each Rad52 focus [i.e. active site] represents a center of recombinational repair capable of processing multiple DNA lesions.”
These are just samples of the exciting findings being made about DNA replication. These and other papers show that it is a fail-safe system with many sophisticated backup and repair mechanisms. While there is still much to learn, and many mysteries to explain, DNA’s ability to replicate is truly a marvel of engineering. Think about the classic chicken-and-egg conundrum for evolution illustrated by (5) above: a gene codes for a protein that is essential for the gene to exist. Browse through the abstracts of these papers just to get a feel for the amazingly complex world of cellular processes going on in your body right now, without your conscious thought or control.
You need energy to make energy
once Earth had pyrophosphite, it had an energetic molecule that, while not as useful as ATP, was at least somewhat similar. “The team found that a compound known as pyrophosphite may have been an important energy source for primitive lifeforms.” Did he have any evidence for this? No; it’s just a requirement. “It’s a chicken and egg question,” he said. “Scientists are in disagreement over what came first – replication, or metabolism. But there is a third part to the equation – and that is energy.” So while scientists are disagreeing about two things, why not add a third? That’s progress: “You need enzymes to make ATP and you need ATP to make enzymes,” explained Dr Kee, as if adding questions qualifies as explaining something: “The question is: where did energy come from before either of these two things existed?” We may not know the answers, but at least our ignorance is getting more sophisticated thanks to OOL research.17
The hardware and software of the cell, evidence of design 18
Paul Davies: the fifth miracle page 62
Due to the organizational structure of systems capable of processing algorithmic (instructional) information, it is not at all clear that a monomolecular system – where a single polymer plays the role of catalyst and informational carrier – is even logically consistent with the organization of information flow in living systems, because there is no possibility of separating information storage from information processing (that being such a distinctive feature of modern life). As such, digital-first systems (as currently posed) represent a rather trivial form of information processing that fails to capture the logical structure of life as we know it. 1
We need to explain the origin of both the hardware and software aspects of life, or the job is only half finished. Explaining the chemical substrate of life and claiming it as a solution to life’s origin is like pointing to silicon and copper as an explanation for the goings-on inside a computer. It is this transition where one should expect to see a chemical system literally take-on “a life of its own”, characterized by informational dynamics which become decoupled from the dictates of local chemistry alone (while of course remaining fully consistent with those dictates). Thus the famed chicken-or-egg problem (a solely hardware issue) is not the true sticking point. Rather, the puzzle lies with something fundamentally different, a problem of causal organization having to do with the separation of informational and mechanical aspects into parallel causal narratives. The real challenge of life’s origin is thus to explain how instructional information control systems emerge naturally and spontaneously from mere molecular dynamics.
Software and hardware are irreducibly complex and interdependent. There is no reason for information processing machinery to exist without the software, and vice versa.
Systems of interconnected software and hardware are irreducibly complex.
All cellular functions are irreducibly complex
Paul Davies, the fifth miracle page 53:
Pluck the DNA from a living cell and it would be stranded, unable to carry out its familiar role. Only within the context of a highly specific molecular milieu will a given molecule play its role in life. To function properly, DNA must be part of a large team, with each molecule executing its assigned task alongside the others in a cooperative manner. Acknowledging the interdependability of the component molecules within a living organism immediately presents us with a stark philosophical puzzle. If everything needs everything else, how did the community of molecules ever arise in the first place? Since most large molecules needed for life are produced only by living organisms, and are not found outside the cell, how did they come to exist originally, without the help of a meddling scientist? Could we seriously expect a Miller-Urey type of soup to make them all at once, given the hit-and-miss nature of its chemistry? 19
Being part of a large team, cooperative manner, inter dependability, everything needs everything else, are just other words for irreducibility and interdependence.
For a nonliving system, questions about irreducible complexity are even more challenging for a totally natural non-design scenario, because natural selection — which is the main mechanism of Darwinian evolution — cannot exist until a system can reproduce. For the origin of life, we can think about the minimal complexity that would be required for reproduction and other basic life functions. Most scientists think this would require hundreds of biomolecular parts, not just the five parts in a simple mousetrap or in my imaginary LMNOP system. And current science has no plausible theories to explain how the minimal complexity required for life (and the beginning of biological natural selection) could have been produced by natural processes before the beginning of biological natural selection.
While the heart serves as a “pump” to deliver blood throughout the body, it also requires oxygenated blood in order to remain healthy. Or consider that the production of red blood cells, one of the major constituents within the vascular system, is regulated by erythropoietin—a hormone produced in the kidney. Yet that kidney requires red blood cells to deliver oxygenated blood. So which evolved first? 22
The Interdependency of Lipid Membranes and Membrane Proteins
Even in the simplest cells, the membrane is a biological device of a staggering complexity that carries diverse protein complexes mediating energy-dependent – and tightly regulated - import and export of metabolites and polymers
Remarkably, even the author of the book: Agents Under Fire: Materialism and the Rationality of Science, pgs. 104-105 (Rowman & Littlefield, 2004). HT: ENV. asks the readers: Hence a chicken and egg paradox: a lipid membrane would be useless without membrane proteins but how could membrane proteins have evolved in the absence of functional membranes?
Biosynthesis of cell membranes: It takes membranes to make membranes
Cell walls indeed provide essential structural support and external interactions in modern organisms (Albers & Meyer, 2011), Despite the stunning diversity that exists among prokaryotic cell envelopes, the synthesis of many of their main components:
1.N-or O-glycosylated S-layer proteins
2.peptidoglycan,
3.O-antigen LPS
4.teichoic acids
5.exopolysaccharides
relies on comparable glycosylation pathways These pathways are all located in the cell membranes, are mediated by similar lipid carriers and have the same orientation across the membrane.
That raises the question: If in extant cells these pathways are located in the cell membranes, where were to location prior membranes existed ? Would it not be correct to say : It takes membranes to make membranes ?
1) https://reasonandscience.catsboard.com/t2117-nuclear-pores#3762
2) http://www.evolutionnews.org/2011/08/direct_rna_templating_a_failed050121.html
3) http://www.fis.puc.cl/~jalfaro/astrobiologia/Astrobiologiavasquez.pdf
4) Meyer, signature of the cell, page 111
5) https://reasonandscience.catsboard.com/t2285-iron-sulfur-clusters-basic-building-blocks-for-life#4646
6) https://reasonandscience.catsboard.com/t2043-dna-error-checking-and-repair#4669
7) https://reasonandscience.catsboard.com/t2043-dna-and-rna-error-checking-and-repair-amazing-evidence-of-design#4671
8 https://reasonandscience.catsboard.com/t1849p30-dna-replication-of-prokaryotes#4672
9) http://www.ideacenter.org/contentmgr/showdetails.php/id/845
10) https://reasonandscience.catsboard.com/t1796-glycolysis?highlight=glycolysis
11) http://www.scientificamerican.com/article/a-simpler-origin-for-life/
12) http://www.uncommondescent.com/intelligent-design/do-viruses-help-explain-the-origin-of-life/
13) https://en.wikipedia.org/wiki/Reverse_transcriptase#In_eukaryotes
14) Fazale Rana, Cell's design, page 186
16) https://reasonandscience.catsboard.com/t1331-the-cell-membrane-irreducible-complexity
17) http://creationsafaris.com/crev201005.htm
18) https://reasonandscience.catsboard.com/t2221-the-hardware-and-software-of-the-cell-evidence-of-design?highlight=software
19) https://reasonandscience.catsboard.com/t2179-the-cell-is-a-interdependent-irreducible-complex-system
20) https://reasonandscience.catsboard.com/t2285-iron-sulfur-clusters-basic-building-blocks-for-life
21) https://reasonandscience.catsboard.com/t2028-origin-of-the-dna-double-helix#3426
22) http://www.apologeticspress.org/ApPubPage.aspx?pub=1&issue=571&article=450
23) https://reasonandscience.catsboard.com/t2371-how-cellular-enzymatic-and-metabolic-networks-point-to-design#5139
24) https://www.quantamagazine.org/lifes-first-molecule-was-protein-not-rna-new-model-suggests-20171102/
Last edited by Otangelo on Sun Aug 07, 2022 1:59 pm; edited 6 times in total