The Nucleolus a Ribosome producing factory
The Nucleolus - a quality control, and storage warehouse for protein recovery - by evolution, or design?
https://reasonandscience.catsboard.com/t3039-the-nucleolus-a-ribosome-producing-factory
For more than 30 years, researchers thought that the nucleolus performed a vital but circumscribed role in the nucleus — manufacturing a specific type of RNA, dubbed rRNA, that assembles into ribosomes, the organelles that make proteins. But scientists have come to realize that the nucleolus is much more complex than rRNA synthesis. Besides serving as a ribosome factory, the organelle also functions as a command center that monitors a cell’s condition and orchestrates responses when it’s under stress. By storing certain proteins and doling them out when they are required, the nucleolus "endows cells with an accurate way to regulate distribution of proteins. Ultimately, the nucleolus helps determine whether cells reproduce and when they die. 1
There are a vast number of protein-RNA-DNA interactions going on in there (involving a very long list of different proteins), so many aspects of nucleolar function are still not known.
At least one function is as an emergency refuge for nuclear proteins during stress. They associated with other proteins to stabilize their conformations and keep them from aggregating, and then when conditions improve, chaperones such as HSP70 help extricate and refold them. The liquid nature of the granular component (GC) region seems to be a key feature in keeping the proteins from further damage, and it only has so much capacity. Under prolonged stress, it apparently gets overloaded, and eventually goes from a liquid state to more of a stiff gel. In this case, that transition seems to be irreversible – HSP70 et al. can’t bring things back, and if the cell survives such treatment at all that the solidified material eventually gets dragged off for some sort of degradation and recycling.
So the nucleolus is a sort of a warehouse for nuclear proteins in times of stress. In this quality control system, they’re preserved in the liquid-phase region in a form that allows them to be refolded later on (or, presumably, to be easily ubiquitinated (a protein is inactivated by attaching ubiquitin to it) for the nuclear proteasomes to demolish them if refolding doesn’t work out). The alternative is for such proteins to aggregate into amyloid-type insoluble forms that can’t be easily refolded or degraded at all, and we’re seeing the machinery that’s grown up around them to keep this from happening. There are at least two hundred proteins that relocate in this fashion, including some that are considered to be mostly cytosolic, which is interesting..2
The nucleolus functions as a phase-separated protein quality control compartment
Cells have quality control mechanisms that operate under normal growth conditions and during stress to maintain protein homeostasis (proteostasis) and prevent the formation of potentially toxic aggregates. Research in recent decades has identified complex quality control systems in the cytoplasm that mediate protein folding, prevent misfolding, and cooperate in protein degradation with the proteasome and autophagy pathways. 3
My comment: Like the multiple error-check and repair mechanisms in the cell, providing a warehouse for protecting proteins in time of stress evidences a clear planned logic, purposeful strategies to economize energy. Recovering proteins from stress is far less energy dispendious than synthesis de-novo. Evolution does not have foresight, and seems to be incapable to "invent" such ingenious and surprising solutions, which hardly someone would ever have predicted on a molecular level !!
The nucleolus is the most obvious structure seen in the nucleus of a eukaryotic cell when viewed in the light microscope. It was so closely scrutinized by early cytologists that an 1898 review could list some 700 references. We now know that the nucleolus is the site for the processing of rRNAs and their assembly into ribosome subunits. Unlike many of the major organelles in the cell, the nucleolus is not bound by a membrane instead, it is a huge aggregate of macromolecules, including the rRNA genes themselves, precursor rRNAs, mature rRNAs, rRNA-processing enzymes, snoRNPs, a large set of assembly factors (including ATPases, GTPases, protein kinases, and RNA helicases), ribosomal proteins, and partly assembled ribosomes.

Electron micrograph of a thin section of a nucleolus in a human fibroblast, showing its three distinct zones.
(A) View of entire nucleus.
(B) Higher-power view of the nucleolus. It is believed that transcription of the rRNA genes takes place between the fibrillar center and the dense fibrillar component and that processing of the rRNAs and their assembly into the two subunits of the ribosome proceeds outward from the dense fibrillar component to the surrounding granular components.
The close association of all these components allows the assembly of ribosomes to occur rapidly and smoothly. Various types of RNA molecules play a central part in the chemistry and structure of the nucleolus. In present-day cells, the rRNA genes have an important role in forming the nucleolus. In a diploid human cell, the rRNA genes are distributed into 10 clusters, located near the tips of five different chromosome pairs. During interphase, these 10 chromosomes contribute DNA loops (containing the rRNA genes) to the nucleolus; in M phase, when the chromosomes condense, the nucleolus fragments and then disappears. Then, in the telophase part of mitosis, as chromosomes return to their semi-dispersed state, the tips of the 10 chromosomes reform small nucleoli, which progressively coalesce into a single nucleolus
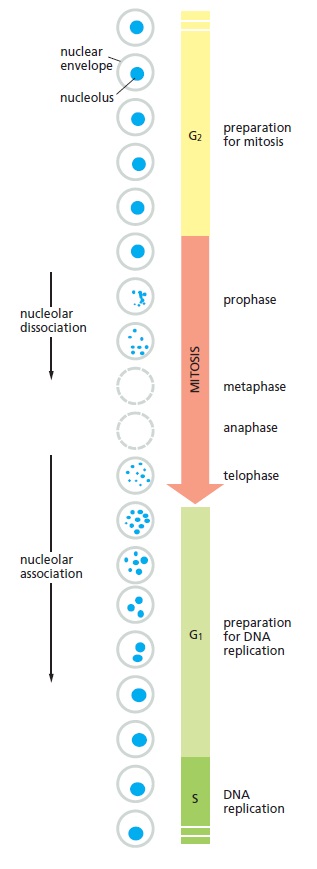
Changes in the appearance of the nucleolus in a human cell during the cell cycle.
Only the cell nucleus is represented in this diagram. In most eukaryotic cells, the nuclear envelope breaks down during mitosis, as indicated by the dashed circles.
The size of the nucleolus reflects the number of ribosomes that the cell is producing. Its size therefore varies greatly in different cells and can change in a single cell, occupying 25% of the total nuclear volume in cells
that are making unusually large amounts of protein. Ribosome assembly is a complex process, the most important features of which are outlined below:
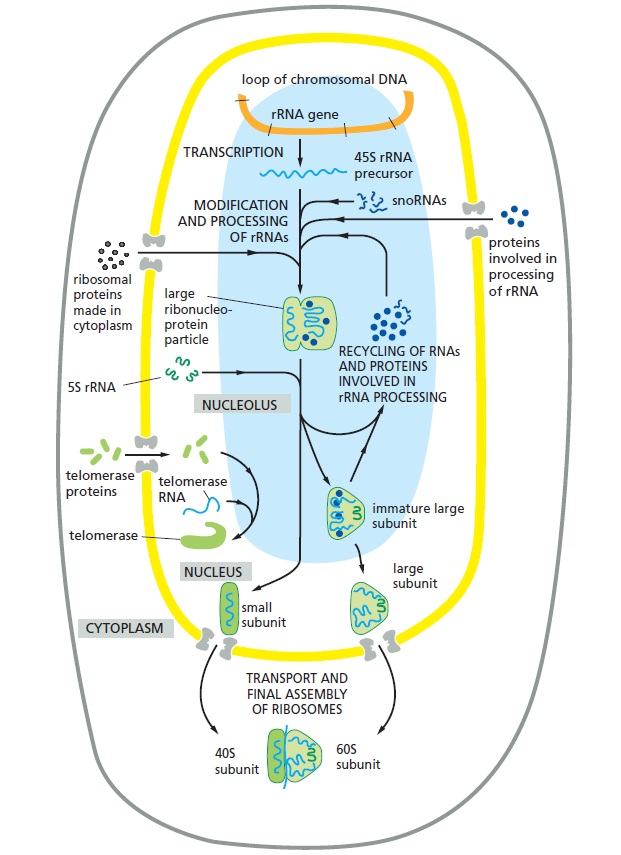
The function of the nucleolus in ribosome and other ribonucleoprotein synthesis.
The 45S precursor rRNA is packaged in a large ribonucleoprotein particle containing many ribosomal proteins imported from the cytoplasm. While this particle remains at the nucleolus, selected components are added and others discarded as it is processed into immature large and small ribosomal subunits. The two ribosomal subunits attain their final functional form only after each is individually transported through the nuclear pores into the cytoplasm. Other ribonucleoprotein complexes, including telomerase shown here, are also assembled in the nucleolus.
My comment: How was this transport programmed and orchestrated? By unguided, nonintelligent processes? Trial and error ?
In addition to its central role in ribosome biogenesis, the nucleolus is the site where other noncoding RNAs are produced and other RNA–protein complexes are assembled. For example, the U6 snRNP, which functions
in pre-mRNA splicing, is composed of one RNA molecule and at least seven proteins. The U6 snRNA is chemically modified by snoRNAs in the nucleolus before its final assembly there into the U6 snRNP. Other important
RNA–protein complexes, including telomerase and the signal-recognition particle , are assembled at the nucleolus. Finally, the tRNAs (transfer RNAs) that carry the amino acids for protein synthesis are processed there as well; like the rRNA genes, the genes encoding tRNAs are clustered in the nucleolus. Thus, the nucleolus can be thought of as a large factory at which different noncoding RNAs are transcribed, processed, and
assembled with proteins to form a large variety of ribonucleoprotein complexes.
The Nucleus Contains a Variety of Subnuclear Aggregates
Although the nucleolus is the most prominent structure in the nucleus, several other nuclear bodies have been observed and studied These include Cajal bodies (named for the scientist who first described them in 1906)
and interchromatin granule clusters (also called “speckles”). Like the nucleolus, these other nuclear structures lack membranes and are highly dynamic depending on the needs of the cell. Their assembly is likely mediated by the association of low complexity protein domains. Their appearance is the result of the tight association of protein and RNA components involved in the synthesis, assembly, and storage of macromolecules involved in gene expression. Cajal bodies are sites where the snRNPs and snoRNPs undergo their final maturation steps, and where the snRNPs are recycled and their RNAs are “reset” after the rearrangements that occur during splicing. In contrast, the interchromatin granule clusters have been proposed to be stockpiles of fully mature snRNPs and other RNA processing components that are ready to be used in the production of mRNA.
Scientists have had difficulties in working out the function of these small subnuclear structures, in part because their appearances can change dramatically as cells traverse the cell cycle or respond to changes in their environment. Moreover, disrupting a particular type of nuclear body often has little effect on cell viability. It seems that the main function of these aggregates is to bring components together at high concentration in order to speed up their assembly. For example, it is estimated that assembly of the U4/U6 snRNP occurs ten times more rapidly in Cajal bodies than would be the case if the same number of components were dispersed throughout the nucleus. Consequently, Cajal bodies appear dispensible in many types of cells but are absolutely required in situations where cells must proliferate rapidly, such as in early vertebrate development.
Here, protein synthesis (which depends on RNA splicing) must be especially rapid, and delays can be lethal. Given the prominence of nuclear bodies in RNA processing, it might be expected that pre-mRNA splicing would occur in a particular location in the nucleus, as it requires numerous RNA and protein components. However, as we have seen, the assembly of splicing components on pre-mRNA is co-transcriptional; thus, splicing must occur at many locations along chromosomes. Although a typical mammalian cell may be expressing on the order of 15,000 genes, transcription and RNA splicing takes place in only several thousand sites in the
nucleus. These sites are highly dynamic and probably result from the association of transcription and splicing components to create small factories, the name given to specific aggregates containing a high local concentration of selected components that create biochemical assembly lines
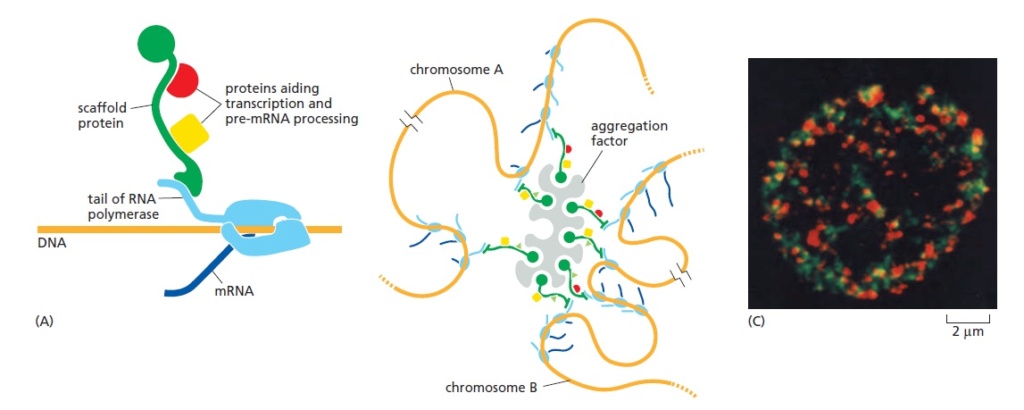
A model for an mRNA production factory.
mRNA production is made more efficient in the nucleus by an aggregation of the many components needed for transcription and pre-mRNA processing, thereby producing a specialized biochemical factory. In (A), a postulated scaffold protein holds various components in the proximity of a transcribing RNA polymerase. Other key components are bound directly to the RNA polymerase tail, which likewise serves as a scaffold, but for simplicity these are not shown here. In (B), a large number of such scaffolds have been brought together to form an aggregate that is highly enriched in the many components needed for the synthesis and processing of pre-mRNAs. Such a scaffold model can account for the several thousand sites of active RNA transcription and processing typically observed in the nucleus of a mammalian cell, each of which has a diameter of roughly 100nm and is estimated to contain, on average, about 10 RNA polymerase II molecules in addition to many other proteins. (C) Here, mRNA production factories and DNA replication factories have been visualized in the same mammalian cell by briefly incorporating differently modified nucleotides into each nucleic acid and detecting the RNA and DNA produced using antibodies, one (green) detecting the newly synthesized DNA and the other (red) detecting the newly synthesized RNA.
Cells have complex quality control mechanisms that operate under normal growth conditions and during stress to maintain protein homeostasis (proteostasis) and prevent the formation of potentially toxic aggregates . Subcellular compartments are equipped with specialized stress re-sponse pathways and vary in stress vulnerability. The nuclear proteome is enriched in proteins contain-ing intrinsically disordered or low complexity sequences. These metastable proteins do not populate a thermodynamically stable folded state and tend to aggregate upon conformational stress. Indeed, various neurodegenerative disorders associated with protein aggregation, such as amyotrophic lateral sclerosis (ALS) and Huntington’s disease, are characterized by the presence of intranuclear inclusions. The nucleus contains several non-membrane bound subcompartments. The largest of these is the nucleolus which consists of liquid-like phases that do not intermix, giving rise to distinct zones (Fig. 1A and fig. S1A and B) (22).
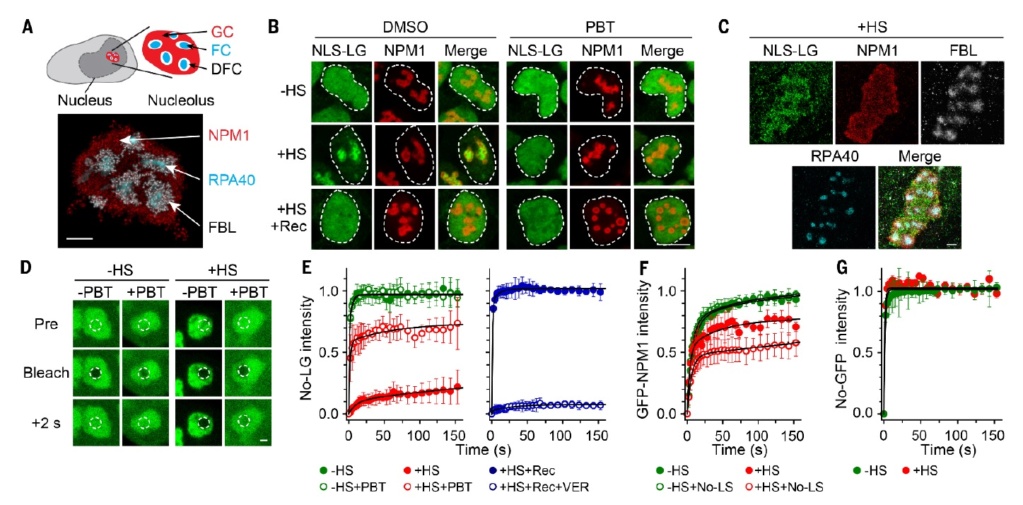
Misfolded proteins transiently accumulate in the GC phase of the nucleolus during stress.
(A) Schematic representation and 3D rendered DNA-PAINT superresolution image of a HeLa cell nucleolus under normal growth condition. NPM1 (GC, red), FBL (DFC, white) and RPA40 (FC, cyan). See also fig. S1, A and B
(B) HEK293T cells stably expressing NLS-LG were treated with DMSO (mock) or PBT before 2 hours of heat stress (+HS), followed by recovery for 2 hours (+HS +Rec). Control cells were maintained at 37°C (-HS). Cells were stained for endogenous NPM1 (red), nuclei are marked by a dashed circle.
(C) Superresolution imaging of HEK293T cells expressing NLS-LG after HS treatment, with staining for GFP, endogenous NPM1, FBL and RPA40. See fig. S1D for –HS control.
(D) No-LG in the nucleolus without (–HS) and with (+HS) heat stress in presence or absence of PBT. Before bleaching (Pre), immediately after bleaching (Bleach) and 2 s after bleaching. (E to G) FRAP analysis of No-LG (E), GFP-NPM1 (F) and No-GFP
(G). No-LG experiments (E) with PBT treatment (empty circles) or DMSO (full circles) as a control. GFP-NPM1 experiments
(F) with cotransfection of No-LS where indicated. For the -HS condition (green), cells were maintained at 37°C during acquisition. For +HS experiments (red), cells were incubated at 43°C for 1 hour before acquisition and maintained at 43°C during acquisition. For the No-LG recovery experiment (E, right graph, blue), cells were subjected to HS and 1 hour recovery (+HS +Rec; full circles), followed by FRAP. Hsp70 was inhibited with VER-155008 before shifting cells to recovery (+HS +Rec +VER; empty circles). CHX was present during recovery. The graphs display corrected and normalized FRAP curves with double exponential fits. Curves are given as mean values ± SD; n ≥ 4 biological repeats representing at least 12 different cells. The first 150 s after photo-bleaching are shown. Quantification of No-LG and GFP-NPM1 mobility is shown in fig. S3, A and B respectively. Scale bars (A, C, D) 1 μm, (B) 10 μm.
Embedded in the outer granular component (GC) phase is the fibrillar center (FC) for the transcription of ribosomal RNA (RNA polymerase I subunit RPA40 as marker). The FC is surrounded by the dense fibrillar component (DFC), which contains the ribonucleoprotein fibrillarin (FBL) (Fig. 1A and fig. S1, A and B). The GC phase is rich in negatively-charged proteins such as nucleophosmin (NPM1) and nucleolin. NPM1 contains extensive unstructured regions and undergoes liquid-liquid phase separation in vitro. During stress, Hsp70 and other molecular chaperones ac-cumulate in the nucleolus, presumably to protect unassembled ribosomal proteins against aggregation. Stress-induced transfer of a nuclear model protein to the nucleolus has also been observed. Here we found that during stress, misfolded proteins entered the liquid-like GC phase of the nucleolus, where irreversible coaggregation of different misfolded protein species was prevented, allowing Hsp70-mediated extraction and refolding (or degradation) upon recovery from stress. In contrast, disruption of the GC phase caused the formation of stable protein aggregates. Prolonged stress resulted in a transition of the nucleolar matrix from liquid-like to solid and prevented quality control.
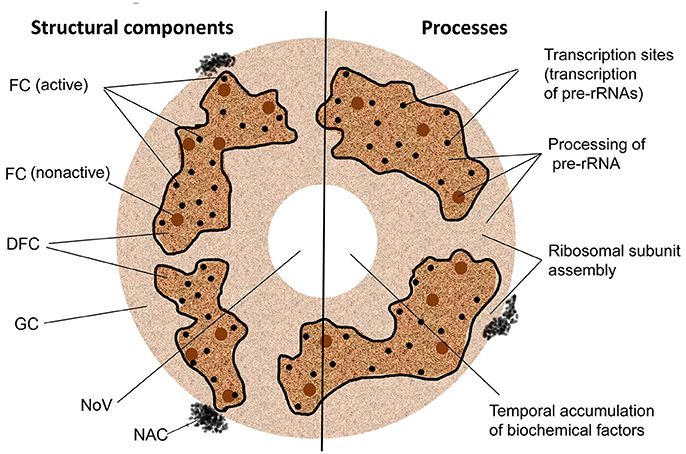
Structural and functional domains of the nucleolus. FC, fibrillar center; DFC, the dense fibrillar component; GC, the granular component; NoV, nucleolar vacuole; NAC, nucleolus-associated chromatin. Nucleolonema is encircled with black lines. 4
The nucleolus is the most conspicuous domain in the eukaryotic cell nucleus, whose main function is ribosomal RNA (rRNA) synthesis and ribosome biogenesis. However, there is growing evidence that the nucleolus is also implicated in many other aspects of cell biology, such as regulation of cell cycle, growth and development, senescence, telomerase activity, gene silencing, responses to biotic and abiotic stresses.
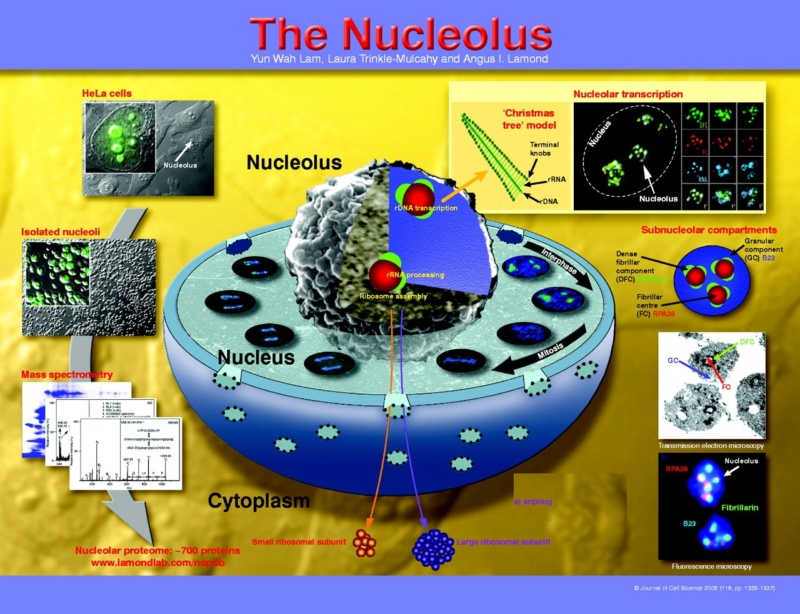
1. https://evolutionnews.org/2014/08/in_the_human_ge_1/
2. https://blogs.sciencemag.org/pipeline/archives/2019/07/24/quality-control-in-the-nucleolus
3. https://science.sciencemag.org/content/365/6451/342
4. https://www.frontiersin.org/articles/10.3389/fpls.2018.00132/full
The Nucleolus - a quality control, and storage warehouse for protein recovery - by evolution, or design?
https://reasonandscience.catsboard.com/t3039-the-nucleolus-a-ribosome-producing-factory
For more than 30 years, researchers thought that the nucleolus performed a vital but circumscribed role in the nucleus — manufacturing a specific type of RNA, dubbed rRNA, that assembles into ribosomes, the organelles that make proteins. But scientists have come to realize that the nucleolus is much more complex than rRNA synthesis. Besides serving as a ribosome factory, the organelle also functions as a command center that monitors a cell’s condition and orchestrates responses when it’s under stress. By storing certain proteins and doling them out when they are required, the nucleolus "endows cells with an accurate way to regulate distribution of proteins. Ultimately, the nucleolus helps determine whether cells reproduce and when they die. 1
There are a vast number of protein-RNA-DNA interactions going on in there (involving a very long list of different proteins), so many aspects of nucleolar function are still not known.
At least one function is as an emergency refuge for nuclear proteins during stress. They associated with other proteins to stabilize their conformations and keep them from aggregating, and then when conditions improve, chaperones such as HSP70 help extricate and refold them. The liquid nature of the granular component (GC) region seems to be a key feature in keeping the proteins from further damage, and it only has so much capacity. Under prolonged stress, it apparently gets overloaded, and eventually goes from a liquid state to more of a stiff gel. In this case, that transition seems to be irreversible – HSP70 et al. can’t bring things back, and if the cell survives such treatment at all that the solidified material eventually gets dragged off for some sort of degradation and recycling.
So the nucleolus is a sort of a warehouse for nuclear proteins in times of stress. In this quality control system, they’re preserved in the liquid-phase region in a form that allows them to be refolded later on (or, presumably, to be easily ubiquitinated (a protein is inactivated by attaching ubiquitin to it) for the nuclear proteasomes to demolish them if refolding doesn’t work out). The alternative is for such proteins to aggregate into amyloid-type insoluble forms that can’t be easily refolded or degraded at all, and we’re seeing the machinery that’s grown up around them to keep this from happening. There are at least two hundred proteins that relocate in this fashion, including some that are considered to be mostly cytosolic, which is interesting..2
The nucleolus functions as a phase-separated protein quality control compartment
Cells have quality control mechanisms that operate under normal growth conditions and during stress to maintain protein homeostasis (proteostasis) and prevent the formation of potentially toxic aggregates. Research in recent decades has identified complex quality control systems in the cytoplasm that mediate protein folding, prevent misfolding, and cooperate in protein degradation with the proteasome and autophagy pathways. 3
My comment: Like the multiple error-check and repair mechanisms in the cell, providing a warehouse for protecting proteins in time of stress evidences a clear planned logic, purposeful strategies to economize energy. Recovering proteins from stress is far less energy dispendious than synthesis de-novo. Evolution does not have foresight, and seems to be incapable to "invent" such ingenious and surprising solutions, which hardly someone would ever have predicted on a molecular level !!
The nucleolus is the most obvious structure seen in the nucleus of a eukaryotic cell when viewed in the light microscope. It was so closely scrutinized by early cytologists that an 1898 review could list some 700 references. We now know that the nucleolus is the site for the processing of rRNAs and their assembly into ribosome subunits. Unlike many of the major organelles in the cell, the nucleolus is not bound by a membrane instead, it is a huge aggregate of macromolecules, including the rRNA genes themselves, precursor rRNAs, mature rRNAs, rRNA-processing enzymes, snoRNPs, a large set of assembly factors (including ATPases, GTPases, protein kinases, and RNA helicases), ribosomal proteins, and partly assembled ribosomes.

Electron micrograph of a thin section of a nucleolus in a human fibroblast, showing its three distinct zones.
(A) View of entire nucleus.
(B) Higher-power view of the nucleolus. It is believed that transcription of the rRNA genes takes place between the fibrillar center and the dense fibrillar component and that processing of the rRNAs and their assembly into the two subunits of the ribosome proceeds outward from the dense fibrillar component to the surrounding granular components.
The close association of all these components allows the assembly of ribosomes to occur rapidly and smoothly. Various types of RNA molecules play a central part in the chemistry and structure of the nucleolus. In present-day cells, the rRNA genes have an important role in forming the nucleolus. In a diploid human cell, the rRNA genes are distributed into 10 clusters, located near the tips of five different chromosome pairs. During interphase, these 10 chromosomes contribute DNA loops (containing the rRNA genes) to the nucleolus; in M phase, when the chromosomes condense, the nucleolus fragments and then disappears. Then, in the telophase part of mitosis, as chromosomes return to their semi-dispersed state, the tips of the 10 chromosomes reform small nucleoli, which progressively coalesce into a single nucleolus

Changes in the appearance of the nucleolus in a human cell during the cell cycle.
Only the cell nucleus is represented in this diagram. In most eukaryotic cells, the nuclear envelope breaks down during mitosis, as indicated by the dashed circles.
The size of the nucleolus reflects the number of ribosomes that the cell is producing. Its size therefore varies greatly in different cells and can change in a single cell, occupying 25% of the total nuclear volume in cells
that are making unusually large amounts of protein. Ribosome assembly is a complex process, the most important features of which are outlined below:

The function of the nucleolus in ribosome and other ribonucleoprotein synthesis.
The 45S precursor rRNA is packaged in a large ribonucleoprotein particle containing many ribosomal proteins imported from the cytoplasm. While this particle remains at the nucleolus, selected components are added and others discarded as it is processed into immature large and small ribosomal subunits. The two ribosomal subunits attain their final functional form only after each is individually transported through the nuclear pores into the cytoplasm. Other ribonucleoprotein complexes, including telomerase shown here, are also assembled in the nucleolus.
My comment: How was this transport programmed and orchestrated? By unguided, nonintelligent processes? Trial and error ?
In addition to its central role in ribosome biogenesis, the nucleolus is the site where other noncoding RNAs are produced and other RNA–protein complexes are assembled. For example, the U6 snRNP, which functions
in pre-mRNA splicing, is composed of one RNA molecule and at least seven proteins. The U6 snRNA is chemically modified by snoRNAs in the nucleolus before its final assembly there into the U6 snRNP. Other important
RNA–protein complexes, including telomerase and the signal-recognition particle , are assembled at the nucleolus. Finally, the tRNAs (transfer RNAs) that carry the amino acids for protein synthesis are processed there as well; like the rRNA genes, the genes encoding tRNAs are clustered in the nucleolus. Thus, the nucleolus can be thought of as a large factory at which different noncoding RNAs are transcribed, processed, and
assembled with proteins to form a large variety of ribonucleoprotein complexes.
The Nucleus Contains a Variety of Subnuclear Aggregates
Although the nucleolus is the most prominent structure in the nucleus, several other nuclear bodies have been observed and studied These include Cajal bodies (named for the scientist who first described them in 1906)
and interchromatin granule clusters (also called “speckles”). Like the nucleolus, these other nuclear structures lack membranes and are highly dynamic depending on the needs of the cell. Their assembly is likely mediated by the association of low complexity protein domains. Their appearance is the result of the tight association of protein and RNA components involved in the synthesis, assembly, and storage of macromolecules involved in gene expression. Cajal bodies are sites where the snRNPs and snoRNPs undergo their final maturation steps, and where the snRNPs are recycled and their RNAs are “reset” after the rearrangements that occur during splicing. In contrast, the interchromatin granule clusters have been proposed to be stockpiles of fully mature snRNPs and other RNA processing components that are ready to be used in the production of mRNA.
Scientists have had difficulties in working out the function of these small subnuclear structures, in part because their appearances can change dramatically as cells traverse the cell cycle or respond to changes in their environment. Moreover, disrupting a particular type of nuclear body often has little effect on cell viability. It seems that the main function of these aggregates is to bring components together at high concentration in order to speed up their assembly. For example, it is estimated that assembly of the U4/U6 snRNP occurs ten times more rapidly in Cajal bodies than would be the case if the same number of components were dispersed throughout the nucleus. Consequently, Cajal bodies appear dispensible in many types of cells but are absolutely required in situations where cells must proliferate rapidly, such as in early vertebrate development.
Here, protein synthesis (which depends on RNA splicing) must be especially rapid, and delays can be lethal. Given the prominence of nuclear bodies in RNA processing, it might be expected that pre-mRNA splicing would occur in a particular location in the nucleus, as it requires numerous RNA and protein components. However, as we have seen, the assembly of splicing components on pre-mRNA is co-transcriptional; thus, splicing must occur at many locations along chromosomes. Although a typical mammalian cell may be expressing on the order of 15,000 genes, transcription and RNA splicing takes place in only several thousand sites in the
nucleus. These sites are highly dynamic and probably result from the association of transcription and splicing components to create small factories, the name given to specific aggregates containing a high local concentration of selected components that create biochemical assembly lines

A model for an mRNA production factory.
mRNA production is made more efficient in the nucleus by an aggregation of the many components needed for transcription and pre-mRNA processing, thereby producing a specialized biochemical factory. In (A), a postulated scaffold protein holds various components in the proximity of a transcribing RNA polymerase. Other key components are bound directly to the RNA polymerase tail, which likewise serves as a scaffold, but for simplicity these are not shown here. In (B), a large number of such scaffolds have been brought together to form an aggregate that is highly enriched in the many components needed for the synthesis and processing of pre-mRNAs. Such a scaffold model can account for the several thousand sites of active RNA transcription and processing typically observed in the nucleus of a mammalian cell, each of which has a diameter of roughly 100nm and is estimated to contain, on average, about 10 RNA polymerase II molecules in addition to many other proteins. (C) Here, mRNA production factories and DNA replication factories have been visualized in the same mammalian cell by briefly incorporating differently modified nucleotides into each nucleic acid and detecting the RNA and DNA produced using antibodies, one (green) detecting the newly synthesized DNA and the other (red) detecting the newly synthesized RNA.
Cells have complex quality control mechanisms that operate under normal growth conditions and during stress to maintain protein homeostasis (proteostasis) and prevent the formation of potentially toxic aggregates . Subcellular compartments are equipped with specialized stress re-sponse pathways and vary in stress vulnerability. The nuclear proteome is enriched in proteins contain-ing intrinsically disordered or low complexity sequences. These metastable proteins do not populate a thermodynamically stable folded state and tend to aggregate upon conformational stress. Indeed, various neurodegenerative disorders associated with protein aggregation, such as amyotrophic lateral sclerosis (ALS) and Huntington’s disease, are characterized by the presence of intranuclear inclusions. The nucleus contains several non-membrane bound subcompartments. The largest of these is the nucleolus which consists of liquid-like phases that do not intermix, giving rise to distinct zones (Fig. 1A and fig. S1A and B) (22).

Misfolded proteins transiently accumulate in the GC phase of the nucleolus during stress.
(A) Schematic representation and 3D rendered DNA-PAINT superresolution image of a HeLa cell nucleolus under normal growth condition. NPM1 (GC, red), FBL (DFC, white) and RPA40 (FC, cyan). See also fig. S1, A and B
(B) HEK293T cells stably expressing NLS-LG were treated with DMSO (mock) or PBT before 2 hours of heat stress (+HS), followed by recovery for 2 hours (+HS +Rec). Control cells were maintained at 37°C (-HS). Cells were stained for endogenous NPM1 (red), nuclei are marked by a dashed circle.
(C) Superresolution imaging of HEK293T cells expressing NLS-LG after HS treatment, with staining for GFP, endogenous NPM1, FBL and RPA40. See fig. S1D for –HS control.
(D) No-LG in the nucleolus without (–HS) and with (+HS) heat stress in presence or absence of PBT. Before bleaching (Pre), immediately after bleaching (Bleach) and 2 s after bleaching. (E to G) FRAP analysis of No-LG (E), GFP-NPM1 (F) and No-GFP
(G). No-LG experiments (E) with PBT treatment (empty circles) or DMSO (full circles) as a control. GFP-NPM1 experiments
(F) with cotransfection of No-LS where indicated. For the -HS condition (green), cells were maintained at 37°C during acquisition. For +HS experiments (red), cells were incubated at 43°C for 1 hour before acquisition and maintained at 43°C during acquisition. For the No-LG recovery experiment (E, right graph, blue), cells were subjected to HS and 1 hour recovery (+HS +Rec; full circles), followed by FRAP. Hsp70 was inhibited with VER-155008 before shifting cells to recovery (+HS +Rec +VER; empty circles). CHX was present during recovery. The graphs display corrected and normalized FRAP curves with double exponential fits. Curves are given as mean values ± SD; n ≥ 4 biological repeats representing at least 12 different cells. The first 150 s after photo-bleaching are shown. Quantification of No-LG and GFP-NPM1 mobility is shown in fig. S3, A and B respectively. Scale bars (A, C, D) 1 μm, (B) 10 μm.
Embedded in the outer granular component (GC) phase is the fibrillar center (FC) for the transcription of ribosomal RNA (RNA polymerase I subunit RPA40 as marker). The FC is surrounded by the dense fibrillar component (DFC), which contains the ribonucleoprotein fibrillarin (FBL) (Fig. 1A and fig. S1, A and B). The GC phase is rich in negatively-charged proteins such as nucleophosmin (NPM1) and nucleolin. NPM1 contains extensive unstructured regions and undergoes liquid-liquid phase separation in vitro. During stress, Hsp70 and other molecular chaperones ac-cumulate in the nucleolus, presumably to protect unassembled ribosomal proteins against aggregation. Stress-induced transfer of a nuclear model protein to the nucleolus has also been observed. Here we found that during stress, misfolded proteins entered the liquid-like GC phase of the nucleolus, where irreversible coaggregation of different misfolded protein species was prevented, allowing Hsp70-mediated extraction and refolding (or degradation) upon recovery from stress. In contrast, disruption of the GC phase caused the formation of stable protein aggregates. Prolonged stress resulted in a transition of the nucleolar matrix from liquid-like to solid and prevented quality control.

Structural and functional domains of the nucleolus. FC, fibrillar center; DFC, the dense fibrillar component; GC, the granular component; NoV, nucleolar vacuole; NAC, nucleolus-associated chromatin. Nucleolonema is encircled with black lines. 4
The nucleolus is the most conspicuous domain in the eukaryotic cell nucleus, whose main function is ribosomal RNA (rRNA) synthesis and ribosome biogenesis. However, there is growing evidence that the nucleolus is also implicated in many other aspects of cell biology, such as regulation of cell cycle, growth and development, senescence, telomerase activity, gene silencing, responses to biotic and abiotic stresses.

1. https://evolutionnews.org/2014/08/in_the_human_ge_1/
2. https://blogs.sciencemag.org/pipeline/archives/2019/07/24/quality-control-in-the-nucleolus
3. https://science.sciencemag.org/content/365/6451/342
4. https://www.frontiersin.org/articles/10.3389/fpls.2018.00132/full

