RNA Epigenetics 1
DNA isn’t the only decorated nucleic acid in the cell. Modifications to RNA molecules are much more common and are critical for regulating diverse biological processes.
The nucleotide building blocks of RNA contain pyrimidine or purine rings, and each position of these rings can be chemically altered by the addition of various chemical groups. Most frequently, a methyl (–CH3) group is tacked on to the outside of the ring. Other chemical additions such as acetyl, isopentenyl, and threonylcarbamoyl are also found added to RNA bases.
The alphabet of RNA consists of at least 140 alternative nucleotide forms. Epigenetic marks are introduced by enzymes and cofactors known as “writers,” and m6A is no exception. This mark is added to RNA by a large protein complex that includes three well-characterized components: METTL3, METTL14, and WTAP.3,4
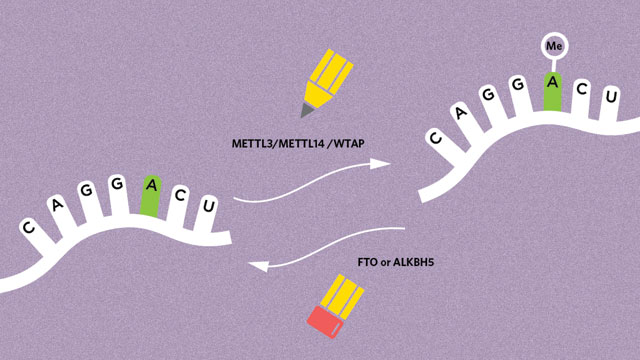
The transcripts of most expressed genes, in a variety of cell types, were shown to be methylated.
Dynamic m6A RNA markings, the new kid on the epigenetic block, herald the era of tripartite epigenetics where modifications of DNA, RNA, and proteins join hands to fine-tune gene expression and to execute prompt and precise responses to environmental stimuli and stresses.
We (G.R. and collaborators) and other groups recently demonstrated that the m6A writer METTL3 is also an essential regulator for termination of mouse embryonic stem cell pluripotency. Knocking out Mettl3 in preimplantation murine epiblasts and in undifferentiated ESCs led to depletion of m6A in mRNAs. Cell viability was not affected, suggesting that m6A decoration is not essential for the maintenance of the ESC naive state, but m6A marks were critical for early differentiation. The loss of this modification led to aberrant and restricted lineage priming at the post-implantation stage, resulting in early embryonic lethality.13 The presence of m6A also decreased mRNA stability, including in those transcripts important for maintaining pluripotency. These findings demonstrated, for the first time, an essential function for an mRNA modification in vivo.14
1. http://mobile.the-scientist.com/article/44873/rna-epigenetics
DNA isn’t the only decorated nucleic acid in the cell. Modifications to RNA molecules are much more common and are critical for regulating diverse biological processes.
The nucleotide building blocks of RNA contain pyrimidine or purine rings, and each position of these rings can be chemically altered by the addition of various chemical groups. Most frequently, a methyl (–CH3) group is tacked on to the outside of the ring. Other chemical additions such as acetyl, isopentenyl, and threonylcarbamoyl are also found added to RNA bases.
The alphabet of RNA consists of at least 140 alternative nucleotide forms. Epigenetic marks are introduced by enzymes and cofactors known as “writers,” and m6A is no exception. This mark is added to RNA by a large protein complex that includes three well-characterized components: METTL3, METTL14, and WTAP.3,4

The transcripts of most expressed genes, in a variety of cell types, were shown to be methylated.
Dynamic m6A RNA markings, the new kid on the epigenetic block, herald the era of tripartite epigenetics where modifications of DNA, RNA, and proteins join hands to fine-tune gene expression and to execute prompt and precise responses to environmental stimuli and stresses.
We (G.R. and collaborators) and other groups recently demonstrated that the m6A writer METTL3 is also an essential regulator for termination of mouse embryonic stem cell pluripotency. Knocking out Mettl3 in preimplantation murine epiblasts and in undifferentiated ESCs led to depletion of m6A in mRNAs. Cell viability was not affected, suggesting that m6A decoration is not essential for the maintenance of the ESC naive state, but m6A marks were critical for early differentiation. The loss of this modification led to aberrant and restricted lineage priming at the post-implantation stage, resulting in early embryonic lethality.13 The presence of m6A also decreased mRNA stability, including in those transcripts important for maintaining pluripotency. These findings demonstrated, for the first time, an essential function for an mRNA modification in vivo.14
1. http://mobile.the-scientist.com/article/44873/rna-epigenetics

