https://reasonandscience.catsboard.com/t2364-the-irreducible-code-instructed-process-to-make-cell-factories-and-machines-points-to-intelligent-design
Would you also say that it is plausible that a tornado over a junkyard could produce a 747?
Would you also say that it is plausible that mindless random chance can write a book?
The cell is a factory, that has various computer like hierarchically organized systems of hardware and software, various language based informational systems, a translation system, hudge amounts of precise instructional/specified, complex information stored and extract systems to make all parts needed to produce the factory and replicate itself, the scaffold structure, that permits the build of the indispensable protection wall, form and size of its building, walls with gates that permits cargo in and out, recognition mechanisms that let only the right cargo in, has specific sites and production lines, "employees", busy and instructed to produce all kind of necessary products, parts and subparts with the right form and size through the right materials, others which mount the parts together in the right order, on the right place, in the right sequence, at the right time, which has sophisticated check and error detection mechanisms all along the production process, the hability to compare correctly produced parts to faulty ones and discard the faulty ones, and repeat the process to make the correct ones; highways and cargo carriers that have tags which recognize where to drop the cargo where it's needed, cleans up waste and has waste bins and sophisticated recycle mechanisms, storage departments, produces its energy and shuttles it to where its needed, and last not least, does reproduce itself.
The salient thing is that the individual parts and compartments have no function by their own. They had to emerge ALL AT ONCE, No stepwise manner is possible, all systems are INTERDEPENDENT and IRREDUCIBLE. And it could not be through evolution, since evolution depends on fully working self-replicating cells, in order to function.
How can someone rationally argue that the origin of the most sophisticated factory in the universe would be probable to be based on natural occurrence, without involving any guiding intelligence?
chemist Wilhelm Huck, professor at Radboud University Nijmegen
A working cell is more than the sum of its parts. "A functioning cell must be entirely correct at once, in all its complexity,"
To go from a bacterium to people is less of a step than to go from a mixture of amino acids to a bacterium. — Lynn Margulis
Evolution has been a central point of the origins debate. Abiogenesis, however, provides far better elucidation of what mechanisms explain the origin of biological systems better: an intelligent designer, through power, information input, wisdom, will, or natural, non-guided, non-intelligent mechanisms, that is: random chance or physical necessity, long periods of time, mutation and natural selection, or self-organisation of matter.
Behe's definition of Irreducible complexity can be expanded and applied not only to biological systems, but also to systems, machines, and factories created by man, that require a minimal number of parts to exercise a specific function, and this minimal number of parts cannot be reduced to keep the basic function. The term applies as well to processes, production methods and proceedings of various sorts. To reach a certain goal, a minimal number of manufacturing steps must be gone through. That applies in special to processes in living cells, where a minimal set of basic processes must be fully functional and operational, in order to maintain cells alive.
Following irreducible processes and parts are required to keep cells alive and illustrate mount improbable to get life a first go:
Reproduction. Reproduction is essential for the survival of all living things.
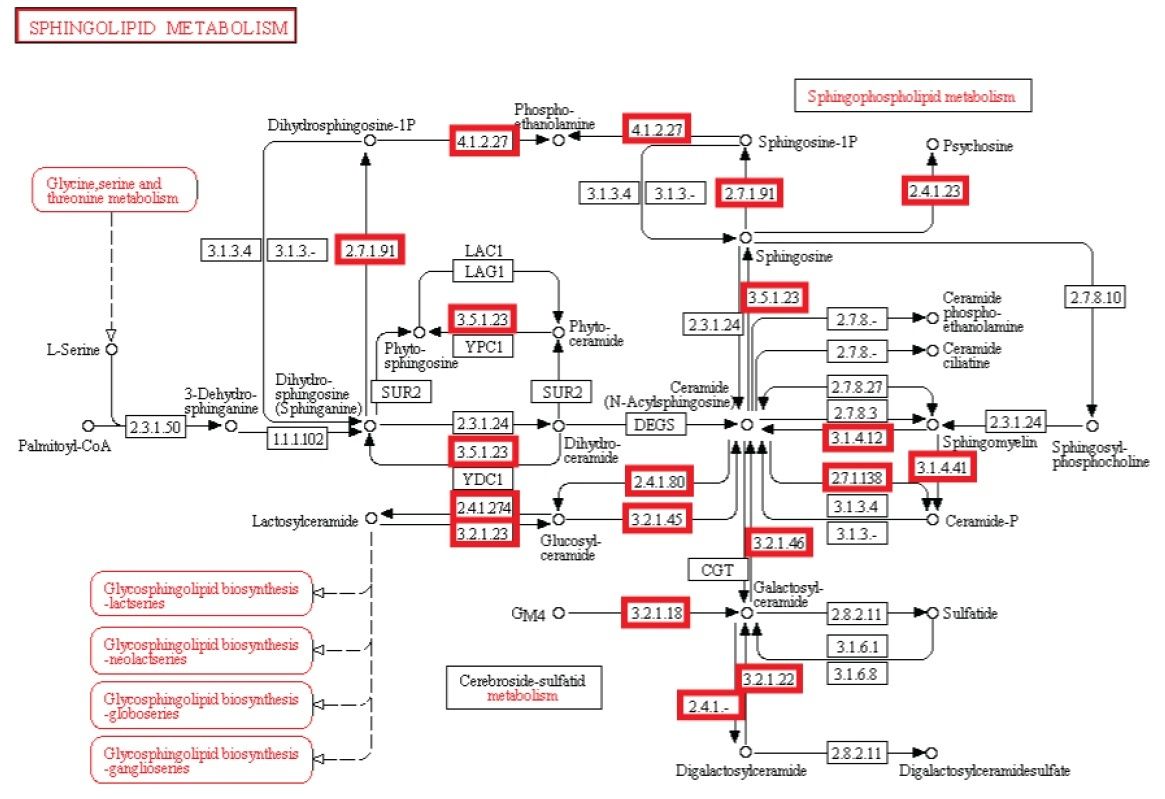
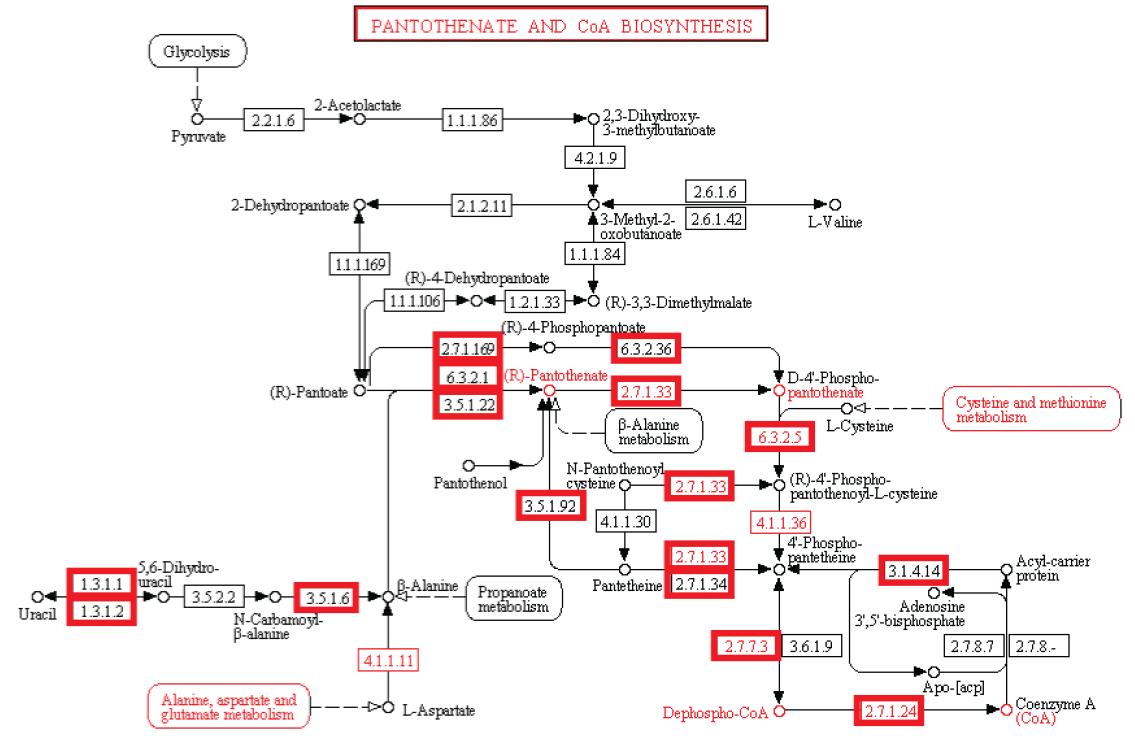
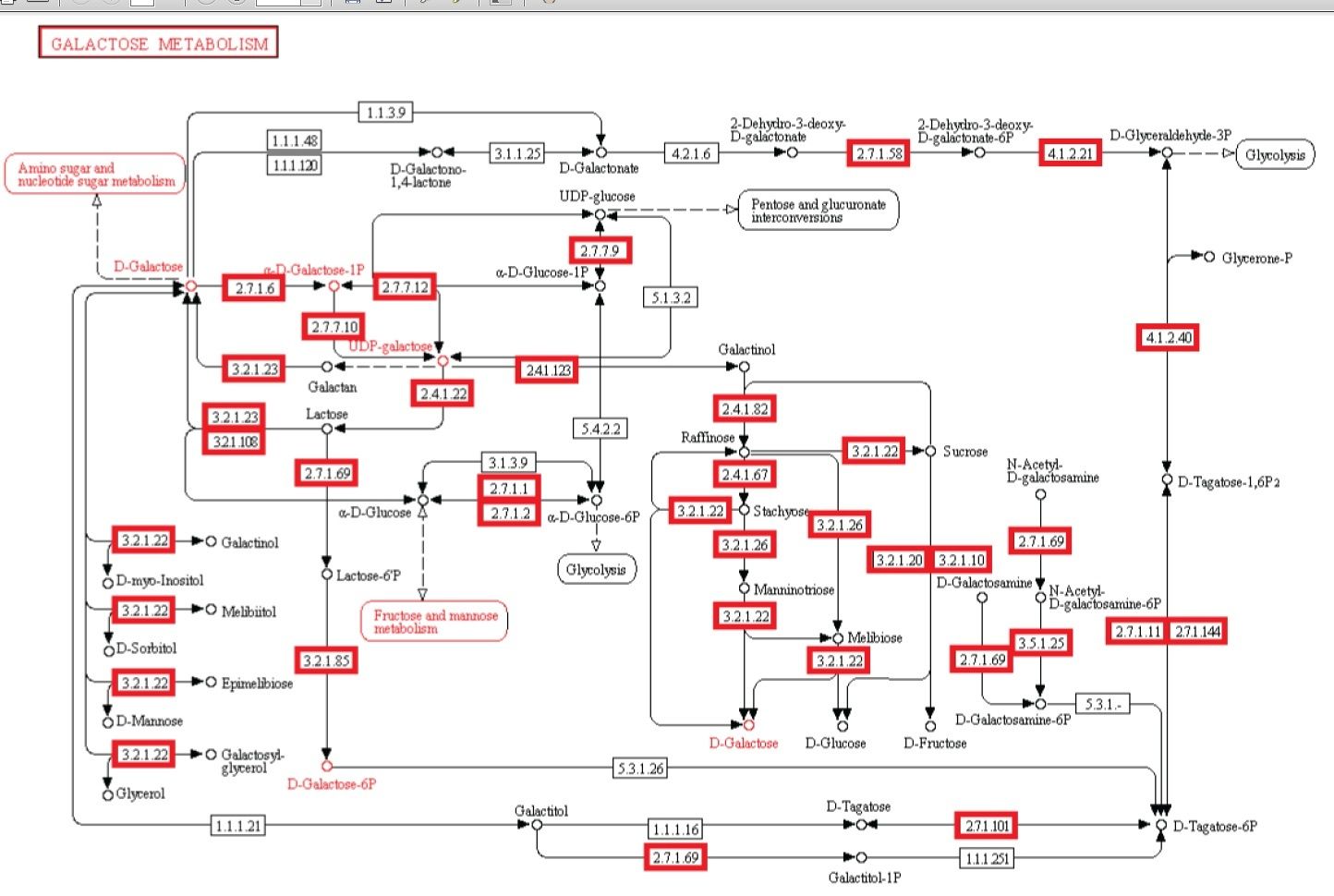
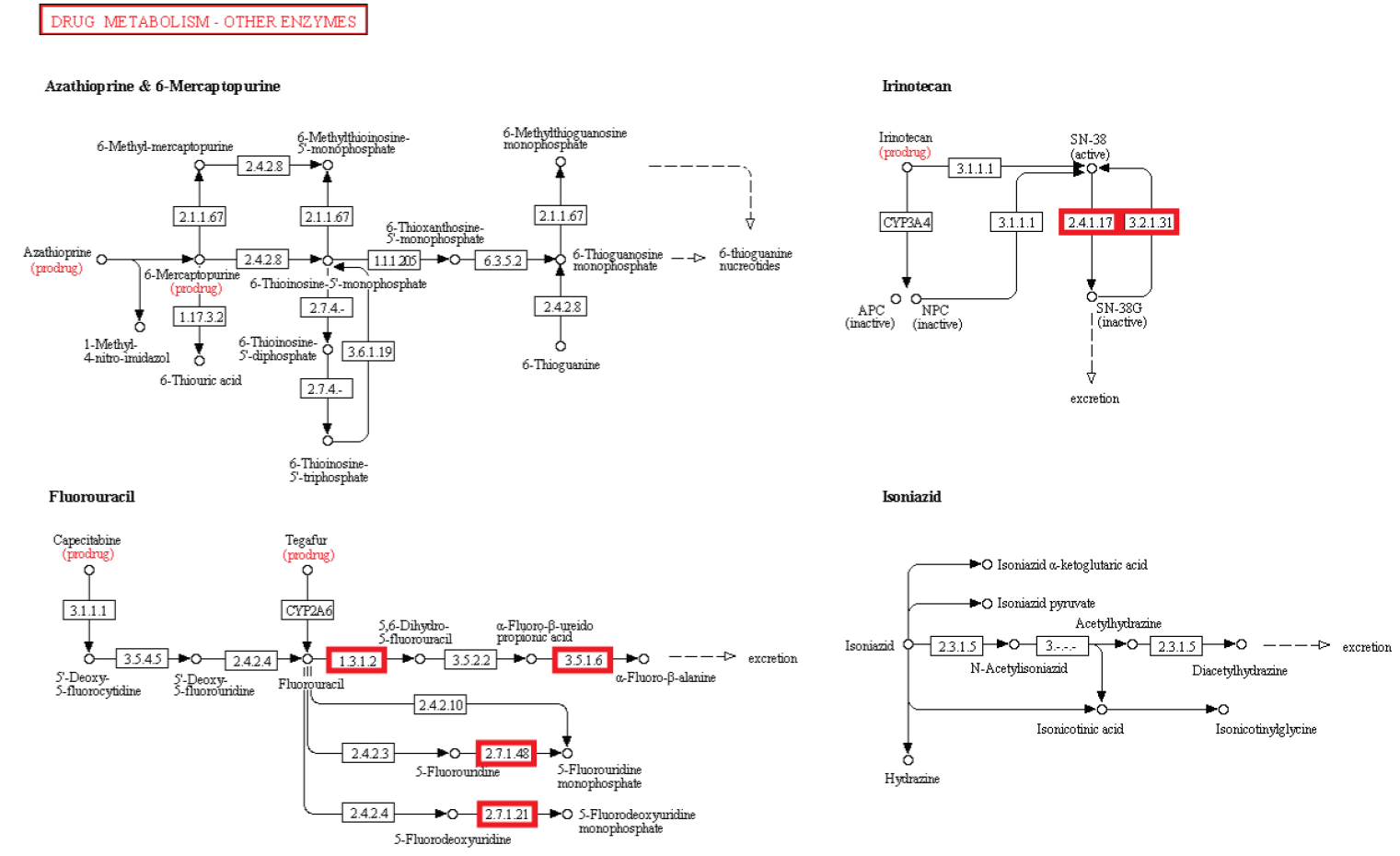
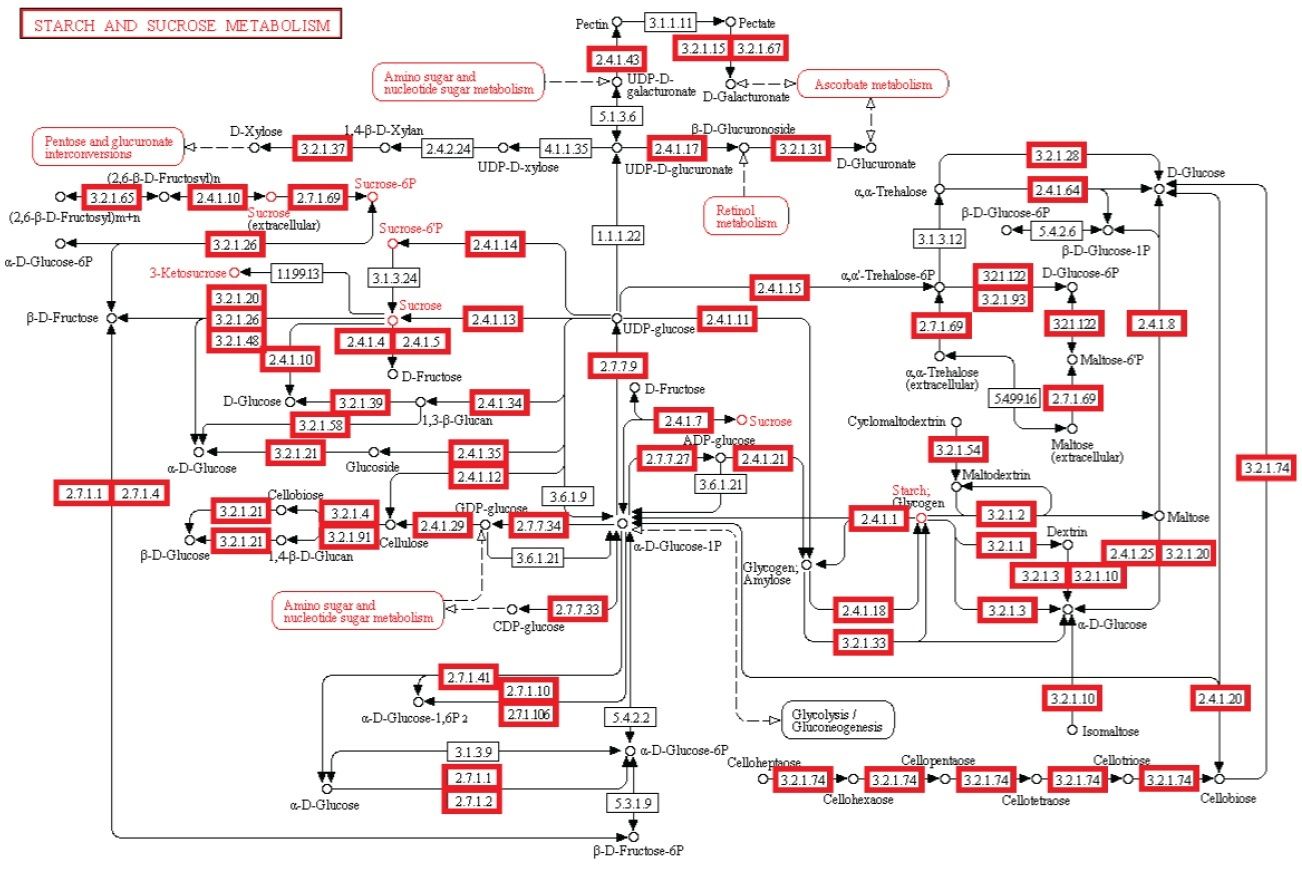
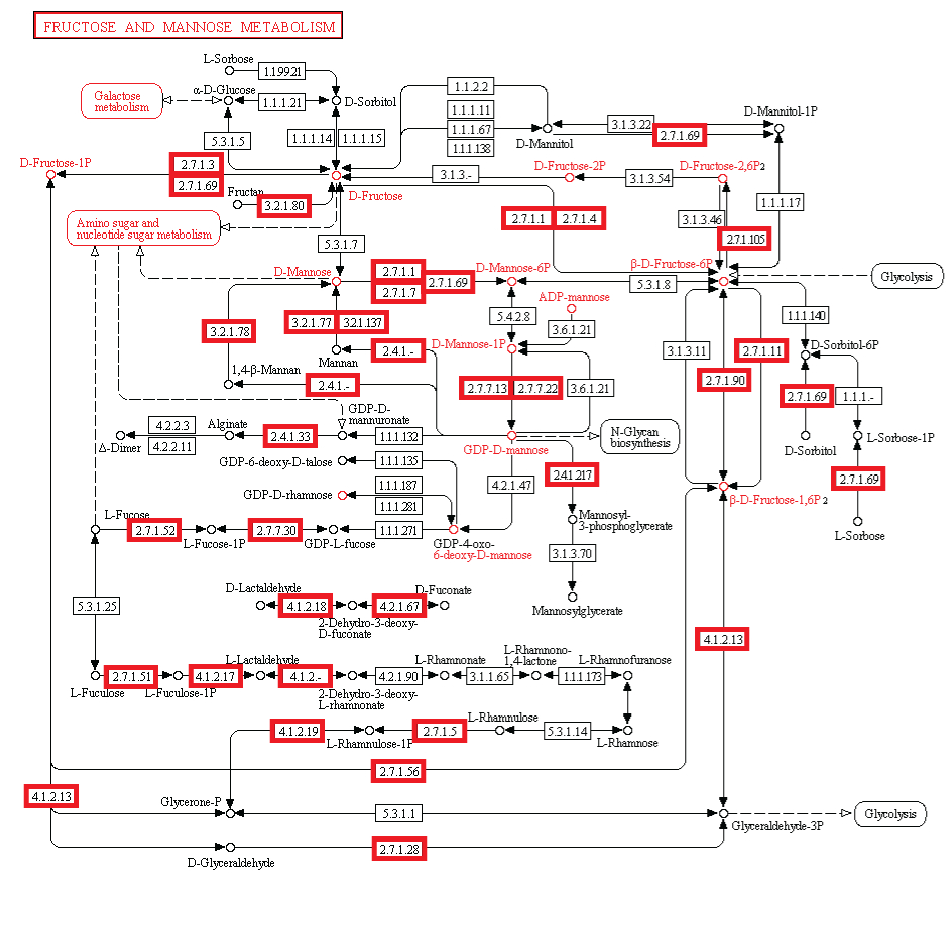
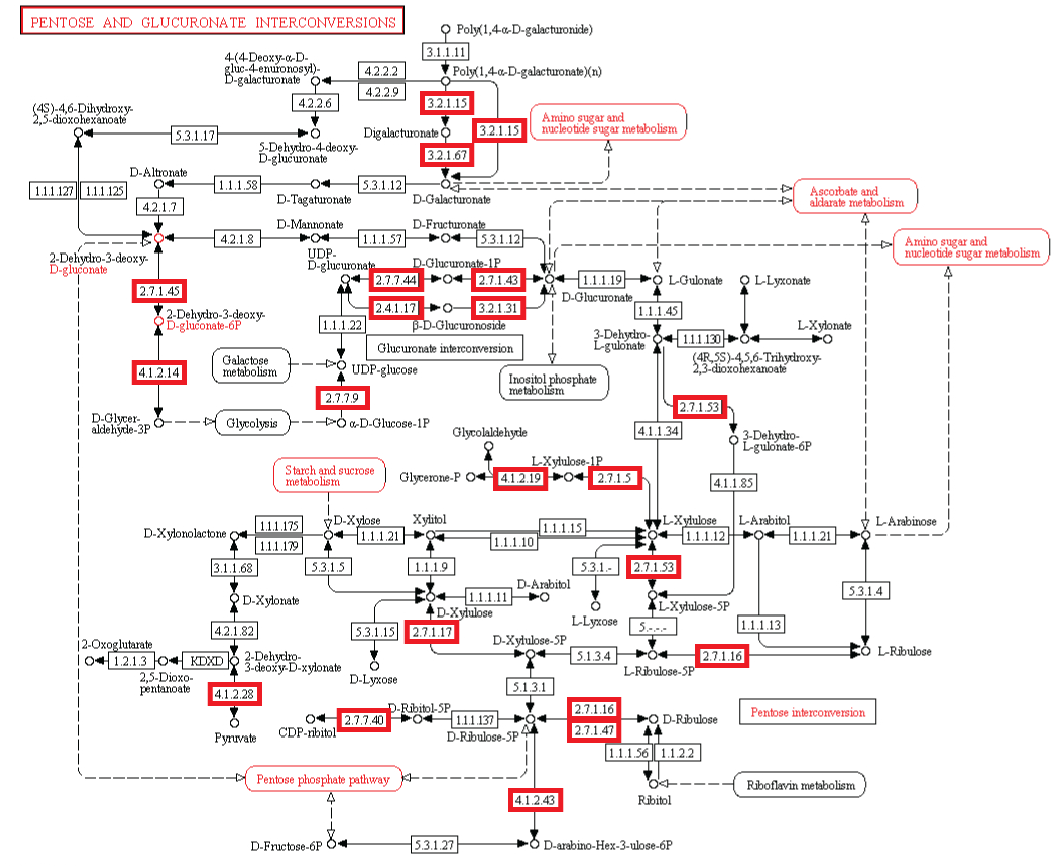
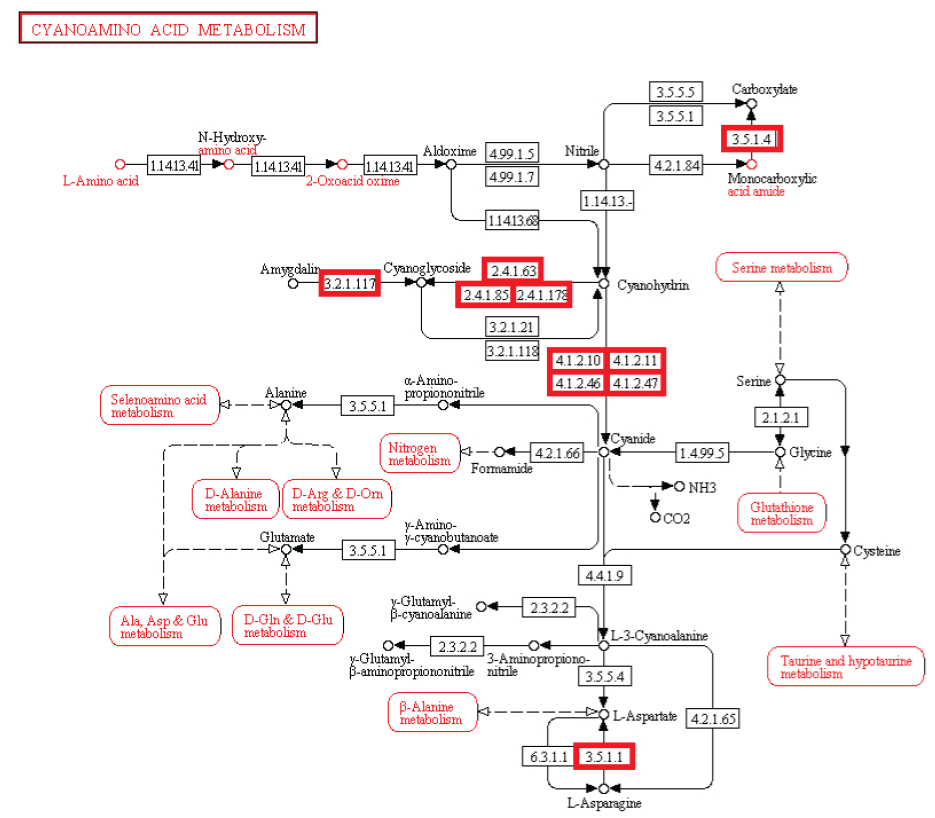
Metabolism. Enzymatic activity allows a cell to respond to changing environmental demands and regulate its metabolic pathways, both of which are essential to cell survival.
Nutrition. This is closely related to metabolism. Seal up a living organism in a box for long enough and in due course, it will cease to function and eventually die. Nutrients are essential for life.
Complexity. All known forms of life are amazingly complex. Even single-celled organisms such as bacteria are veritable beehives of activity involving millions of components.
Organization. Maybe it is not complexity per se that is significant but organized complexity.
Growth and development. Individual organisms grow and ecosystems tend to spread (if conditions are right).
Information content. In recent years scientists have stressed the analogy between living organisms and computers. Crucially, the information needed to replicate an organism is passed on in the genes from parent to offspring.
Hardware/software entanglement. All life of the sort found on Earth stems from a deal struck between two very different classes of molecules: nucleic acids and proteins.
Permanence and change. A further paradox of life concerns the strange conjunction of permanence and change.
Sensitivity. All organisms respond to stimuli— though not always to the same stimuli in the same ways.
Regulation. All organisms have regulatory mechanisms that coordinate internal processes.
Jack W. Szostak The Origins of Cellular Life
We have proposed that a simple primitive cell, or protocell, would consist of two key components: a protocell membrane
that defines a spatially localized compartment and an informational polymer that allows for the replication and inheritance of functional information
Can you believe that?
http://molbio.mgh.harvard.edu/szostakweb/publications/Szostak_pdfs/Schrum_et_al_CSH_2010.pdf
Following is the description of parts and processes in a theoretical protocell, which are essential and irreducible:
What Might Be a Protocell’s minimal requirement of parts?
1. The Cell membrane
2. DNA repair mechanisms
3. Plasma membrane gates
4. The Cytoplasm
5. Glycolysis
6. Left-handed Amino Acids
7. Membrane-enclosed vesicles
8. Ribosomes
9. tRNA
10. right-handed DNA
11. Signal-Recognition Particles (SRP)
12. Lysosomes
13. A complete transcriptional machinery
14. Protein-processing, -folding, secretion, and degradation functions and two proteases.
15. FtsZ
16. Cation, ABC transporters, a PTS for glucose transport, phosphate transporters
17. Dihydroxyacetone phosphate
18. ATP synthase
The Cell membrane separates the interior of all cells from the outside environment. That's the exterior factory wall that protects the factory.
The Nucleus ( only in eukaryotic cells ) is the Chief Executive Officer (CEO). It controls all cell activity; determines what proteins will be made and controls all cell activity.
DNA repair mechanisms Proofreading enzymes’ to prevent the occurrence of slight changes in sequence when DNA replicates.
Plasma membrane gates Control of Input Flow, Functions of Material Identification and Material Extraction. Regulate what enters and leaves the cell; where cells make contact with the external environment. That's the Shipping/Receiving Department. It functions also as the communications department because it is where the cell contacts the external environment.
The Cytoplasm includes everything from the cell membrane and the nucleus. It contains various kinds of cell structures and is the site of most cell activity. The cytoplasm is similar to the factory floor where most of the products are assembled, finished, and shipped.
Membrane-enclosed vesicles form packages for cargo so that they may quickly and efficiently reach their destinations.
Internal membranes divide the cell into specialized compartments, each carrying out a specific function inside the cell. That are the compartments in a manufacturing facility.
Ribosomes build the proteins, equal to the Workers in the assembly line.
Signal-Recognition Particles (SRP) and signal receptors provide variety of instructions informing the cell as to what destination and pathway the protein must follow. That's the address on the parcel where it has to be delivered.
A complete transcriptional machinery. including the three subunits of the RNA polymerase, a factor, an RNA helicase, and four transcriptional factors (with elongation, antitermination, and transcription-translation coupling functions)
Protein-processing, -folding, secretion, and degradation functions. GroEL/S and DnaK/DnaJ/GrpE, the signal recognition particle, its receptor, the three essential components of the translocase channel, and a signal peptidase, one endopeptidase, and two proteases.
FtsZ , essential for Cell division
Cation, ABC transporters, a PTS for glucose transport, phosphate transporters. Substrate transport machinery
Dihydroxyacetone phosphate required for Lipid biosynthesis through phosphatidylethanolamine
ATP synthase
This should make it evident that a theoretical natural, non-intelligence requiring transition from a supposed RNA World to a DNA world, to a fully working living cell, even the most simple, is unlikely to the extreme. It reinforces what Urey and many other scientists, origin of life researchers said : Abiogenesis is impossible.
Harold Urey, a founder of origin-of-life research, describes evolution as a faith which seems to defy logic:
“All of us who study the origin of life find that the more we look into it, the more we feel that it is too complex to have evolved anywhere. We believe as an article of faith that life evolved from dead matter on this planet. It is just that its complexity is so great, it is hard for us to imagine that it did.
Paul Davies, the fifth miracle, page 54:
Life as we know it requires hundreds of thousands of specialist proteins, not to mention the nucleic acids. The odds against producing just the proteins by pure chance are something like 1O^40000 to 1. There are indeed a lot of stars—at least ten billion billion in the observable universe. But this number, gigantic as it may appear to us, is nevertheless trivially small compared with the gigantic odds against the random assembly of even a single protein molecule. Though the universe is big, if life formed solely by random agitation in a molecular junkyard, there is scant chance it has happened twice.
1. High instructional coded complex information content (or specified complexity) and irreducible complexity constitute strong indicators or hallmarks of intervention of a (past) intelligent powerful agent.
2. Cells require high genetic and epigenetic information content (or specified complexity) and utilize systems and subsystems that cannot be reduced.
3. Naturalistic mechanisms or undirected causes do not suffice to explain the origin of information (specified complexity) or irreducible complexity.
4. Therefore, intelligent design constitutes the best explanations for the origin of information and irreducible complexity in biological systems.
Objections:
The biological macromolecules of today are not the same as prebiotic protocells.
Answer :
http://reasonandscience.heavenforum.org/t2176-lucathe-last-universal-common-ancestor#3995
This urancestor accumulated genetic information before the rise of organismal lineages and is considered to be either a simple 'progenote' organism with a rudimentary translational apparatus or a more complex 'cenancestor' with almost all essential biological processes. Recent comparative genomic studies support the latter model and propose that the urancestor was similar to modern organisms in terms of gene content. However, most of these studies were based on molecular sequences, which are fast evolving and of limited value for deep evolutionary explorations.
http://news.illinois.edu/news/11/1005LUCA_ManfredoSeufferheld_JamesWhitfield_Caetano_Anolles.html
New evidence suggests that LUCA was a sophisticated organism after all, with a complex structure recognizable as a cell, researchers report. Their study appears in the journal Biology Direct. The study lends support to a hypothesis that LUCA may have been more complex even than the simplest organisms alive today, said James Whitfield, a professor of entomology at Illinois and a co-author on the study.
A minimal estimate for the gene content of the last universal common ancestor--exobiology from a terrestrial perspective. 2
Using an algorithm for ancestral state inference of gene content, given a large number of extant genome sequences and a phylogenetic tree, we aim to reconstruct the gene content of the last universal common ancestor (LUCA), a hypothetical life form that presumably was the progenitor of the three domains of life. The method allows for gene loss, previously found to be a major factor in shaping gene content, and thus the estimate of LUCA's gene content appears to be substantially higher than that proposed previously, with a typical number of over 1000 gene families, of which more than 90% are also functionally characterized. More precisely, when only prokaryotes are considered, the number varies between 1006 and 1189 gene families while when eukaryotes are also included, this number increases to between 1344 and 1529 families depending on the underlying phylogenetic tree. Therefore, the common belief that the hypothetical genome of LUCA should resemble those of the smallest extant genomes of obligate parasites is not supported by recent advances in computational genomics. Instead, a fairly complex genome similar to those of free-living prokaryotes, with a variety of functional capabilities including metabolic transformation, information processing, membrane/transport proteins and complex regulation, shared between the three domains of life, emerges as the most likely progenitor of life on Earth, with profound repercussions for planetary exploration and exobiology.

Last edited by Admin on Sun Apr 08, 2018 6:20 pm; edited 35 times in total