The bioelectric code: An ancient computational medium for dynamic control of growth and form
 Pattern regulation as a closed-loop homeostatic process. (A)
Pattern regulation as a closed-loop homeostatic process. (A) An egg will give rise to a species-specific anatomical construct. However, DNA does not directly encode the geometrical layout of tissues and organs, requiring a process of decoding the genomic information into the spatial configuration.
(B) This process is usually described as a feed-forward system where the activity of gene-regulatory networks within cells results in the expression of effector proteins that, via structural properties and physical forces, result in the emergence of complex shape. In this view, there is no master plan for pattern − only a bottom-up emergent process driven by self-organization and parallel activity of large numbers of agents (cells); this class of models is difficult to apply to a number of biological phenomena. Some species, including many mammals, utilize regulative development which can adjust to radical deformations to the normal developmental sequence.
(C) Their embryos can be divided in half, giving rise to perfectly normal monozygotic twins each of which has regenerated the missing cell mass.
(D) Their embryos can also be combined, giving rise to a normal (but slightly larger) embryo in which no parts are duplicated.
(E) The ability to achieve a specific target morphology despite different starting configurations (flexible morphogenesis) is clearly revealed in the Xenopus tadpole. The normal deformations of the face from that of a tadpole to that of a frog do not break down when tadpole faces are produced with organs (eyes, nostrils, etc.) in aberrant locations: rather than performing a hardwired set of movements (which would create an abnormal frog face if the components start out from incorrect locations), it orchestrates a set of appropriately altered deformations that cease when the correct frog face is produced. This kind of the pattern-homeostatic process must store a set-point that serves as a stop condition; however, as with most types of memory, it can we specifically modified by experience. In the phenomenon of trophic memory
(F), damage created at a specific point on the branched structure of deer antlers is recalled as ectopic branch points in subsequent years’ antler regeneration. This reveals the ability of cells at the scalp to remember the spatial location of specific damage events and alter cell behavior to adjust morphogenesis appropriately − a pattern memory that stretches across months of time and considerable spatial distance.
(G) These kinds of capabilities suggest that patterning is fundamentally a homeostatic process − a closed-loop control system which employs feedback to minimize the error (distance) between a current shape and the stored target morphology. Although these kinds of decisionmaking models are commonplace in engineering, they are only recently beginning to be employed in biology (Barkai and Ben-Zvi, 2009; Pezzulo and Levin, 2016). Panels A,C were created by Justin Guay of Peregrine Creative. Panel C contains a photo by Oudeschool via Wikimedia Commons. Panels E,F are reprinted with permission from (Vandenberg et al., 2012) and (Bubenik and Pavlansky, 1965) respectively.
It has been recognized since ancient times that the egg of a given species gives rise to an individual with the appropriate anatomy of that species (Figure above).How does this occur? What is responsible for the remarkable multi-scale complexity of metazoan organisms, from the distribution of cell types among tissues to the topological shape and arrangement of the body organs, and the geometric layout of the entire body plan?
It is widely believed that the answer lies within the genome, but it is not that simple; DNA simply encodes specific proteins − there is no direct encoding of the anatomical structure. Thus, it is clear from first principles that pattern control involves a code: the encoding of anatomical positions and structures within the egg or other cell type, and the progressive decoding of this information as cells implement invariant morphogenesis (Fig. B). It should be noted that the current understanding of these codes is in its infancy and many fundamental questions remain to be addressed. Despite the progress of genetics and molecular genomics, we are not yet able to predict the anatomical structure of an organism from its genomic sequence (other than by comparing it to genomes whose anatomy we already know), nor in general, do we know how to encode instructions to cells to induce them to develop anatomical structures to a desired functional specification. Indeed, the mystery is revealed to be even deeper than that of embryogenesis, in which the same initial starting condition (the egg) develops into the appropriate target morphology of a given species. Many types of animals exhibit an extensive capacity for regeneration or remodeling; these organisms can restore complex body organs or appendages after dramatic morphological changes such as amputation. For example, planarian flatworms can rebuild any missing part of their body (including the head), while axolotls can regenerate eyes, limbs, tails, jaws, ovaries, and portions of the brain. Such examples reveal that living systems exhibit highly adaptive and context-sensitive pattern homeostasis. Individual cell behaviors are directed towards the maintenance and repair of a specific anatomical configuration. When the correct target morphology is achieved, large-scale remodeling and growth ceases.
Why do we need a code representing homeostatic goal states in tissue properties?The current paradigm recognizes that
different types of codes participate in pattern control. Examples include gradients of gene products that dictate positional information via chemical signals, such as HOX codes, and epigenetic codes that regulate transcriptional cascades via chromatin modification. However, the processes underlying embryogenesis are largely thought of as a ‘feed-forward’ system: the progressive unrolling of the genome in each cell results in specific cellular events which, integrated over large numbers of cellular agents over space and time, results in the emergence of a complex and highly organized forms. The mainstream consensus is that there is no overall encoding of the target morphology: the process is controlled by local events, and the resulting complex pattern is the result of emergence and self-organization. And yet, many of the examples of complex pattern regulation are challenging to explain as an open-loop, purely-emergent process (Fig. 1C,D). For example, embryos of many species can be cut in half or deformed at early stages and yet, can still achieve the morphology of a normal organism (e.g., monozygotic twins from embryo splitting). The ability to achieve the exact same end result from different starting configurations (e.g., a planarian or salamander limb cut at different positions) is highlighted especially starkly by the process of metamorphosis. Becoming a frog requires the tadpole to rearrange its face − the various craniofacial organs move to new positions during metamorphosis. This is normally a stereotypical process, but it was recently discovered that if “Picasso” tadpoles are created (where the eyes, nostrils, and other structures are in aberrant positions), the animals will still turn into largely normal frogs: the organs move in new ways, but still achieve normal frog face target morphology (Fig. 1E). This means that
genetics does not specify hardwired movements of the organs, but rather contribute to the function of a plastic system that enables diverse responses to abnormal starting states so that an invariant (and thus encoded) outcomes result.
While this kind of pattern memory is clearly stable, itis not read-only − it can be rewritten. Modifications made to the shape and size of limbs in crustacea and amphibians respectively are “learned” by the system, resulting in permanent changes to the target morphology (the pattern towards which regeneration builds) upon future rounds of regeneration. A most impressive example (Fig. 1F) is that of trophic memory in deer, in which some species shed and re-grow a consistent branching pattern of antlers (bone and innervation) each year. It was observed that damage made to one point in the branched structure resulted in ectopic branches being produced at the same point in subsequent years of growth. This means that the growth plate in the scalp somehow ‘remembers the location of damage for months, as the whole antler rack falls off and is regenerated, and then triggers the cell behaviors needed to form an ectopic branch in just the right place.
This type of spatial memory in remaining scalp cells (recalling events that occurred at a significant distance in space and time) is especially difficult to reconcile with typical “molecular pathway” arrow models (or gene-regulatory networks) and strongly suggests a spatial encoding system.
Taken together, these examples strongly suggest a ‘closed-loop’ (feed-back based) pattern homeostatic process (Fig. 1G). Systems guided by pure emergence are notoriously difficult to control and study − knowing which low-level rule to perturb experimentally and how to alter it, in order to reach a desired large-scale outcome in a recurrent process is an extremely difficult inverse problem (imagine trying to determine how to modify a function such as z = z2 + c if one wants to add an extra geometric feature to its resulting fractal image). If modular,
representational information (i.e., the encoding of large-scale structure) exists, then re-writing the code to allow the cells to “build to spec” might enable much more efficient control of growth and form compared to approaches that by micromanage individual cell behaviors. In its flight from vitalism and teleology, modern biology has preferred models of emergence and de-centralized control. However, explicitly represented goal states no longer need to be anathema to biology.
Over the last 50+ years, cybernetics, control systems theory, and computer science have revealed frameworks for rigorous means of implementing mechanisms that store complex states and pursue them as homeostatic setpoints.Here, we argue that it is time to consider the possibility that the known emergent features of cell behavior are augmented by a complementary set of top-down controls in which at least some aspects of target anatomical states are encoded within tissue properties.My comment: Top-down encoding, of course, raises the question: Who or what was on top, doing the encoding ? is that no evidence of intelligent design?If tissues must somehow remember at least some aspect of a target state in their physicochemical properties, then encoding and decoding is necessary, since living tissues are storing information about a future (counterfactual) state in their current structure − this is the quintessential context of code. What mechanisms could underlie such a morphogenetic code? Based on the observed examples of stable yet re-writeable anatomical structure, it would have to be a system that supported long-term but labile memory, with capability to sense/measure large-scale spatio-temporal signals. It would also have to have holographic properties (storing information about the whole in individual pieces) and be able to harness individual cell activities toward group-level goals.
While many architectures could in principle support this kind of control system, two well-studied examples do all of the above: cognition in the living brain, and engineered artificial information-processing devices (computers). What these systems have in common is their reliance on electrical signaling. We next consider the role of bioelectrical events in encoding and decoding anatomical structure, keeping in mind that bioelectrics is perhaps but a single component of the rich morphogenetic code that regulates the shape of life at all scales.
Developmental bioelectricity: an encoding medium for pattern controlNeural and non-neural Brains use electrical networks to implement decision-making and memory and to harness the triggering of specific cell behaviors (muscle contractions, gland secretions, and changes in cell proliferation and gene expression)to large-scale goals that are represented in cognitive constructs. It is becoming increasingly appreciated that communication via electrical processes is not unique to nervous systems. The major ion channel families are present near the origin of multicellularity and show significant expansion with the complexification of the metazoan body plan.
The same is true of neurotransmitter signaling, revealing that modern neural networks are an optimized form of a much more primitive cell type that utilized these same pathways to handle its physiological, behavioral, and structural decision-making.While the somatic bioelectric dynamics in these early forms are still unknown, a large literature on electrophysiology in aneural systems from plants, to fungi, to unicellular algae, to bacteria reveals that computational value of physiological networks exists from very early on. Recognizing the need for a medium that supports complex decision-making, classical biologists investigated developmental bioelectrics at the very dawn of mechanistic investigations into the control of body patterning. More recently, with studies of ion channel expression and function, as well as work on pre-neural neurotransmitter roles (Buznikov et al., 1996; Levin et al., 2006; Sullivan and Levin, 2016), the parallels between brain and body in terms of electrical dynamics are becoming ever clearer (Fig. A below).
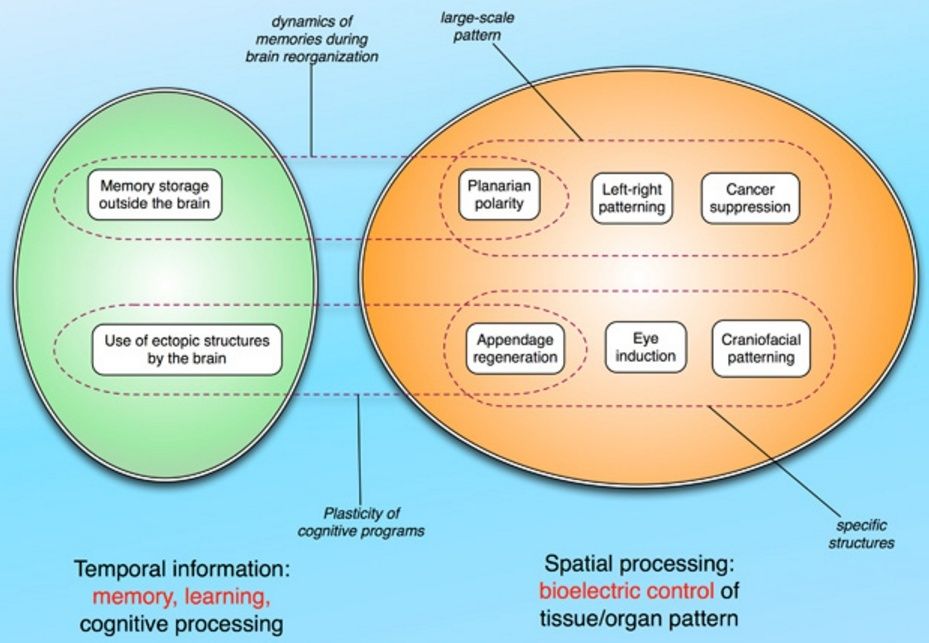
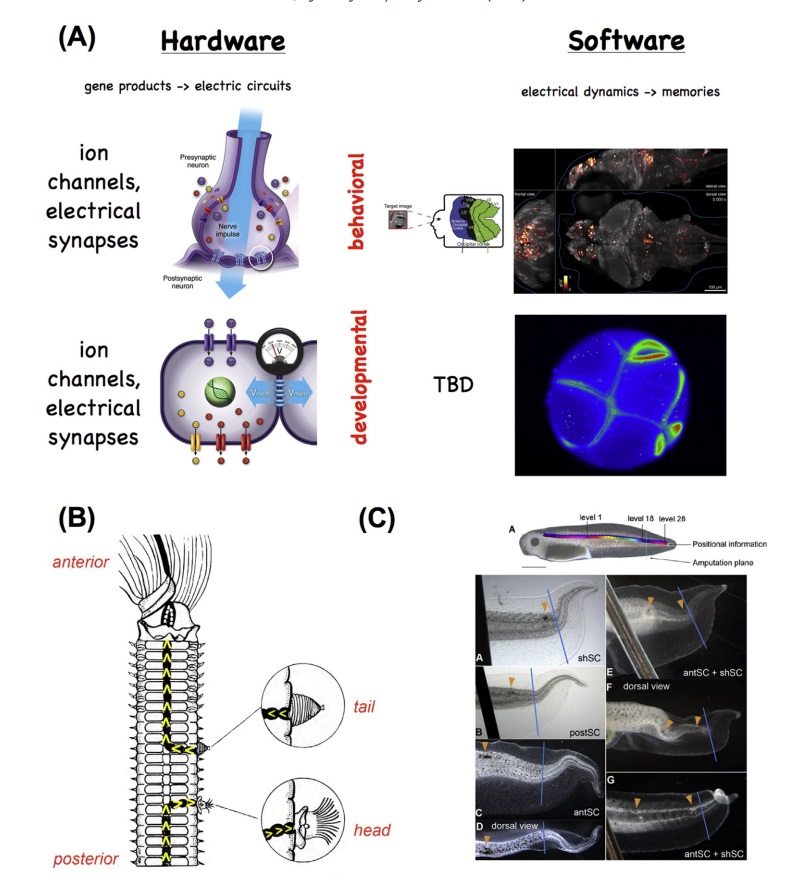 Mechanisms and functionality conserved between brain function and pattern regulation.
Mechanisms and functionality conserved between brain function and pattern regulation. (A) The hardware of the brain consists of ion channels that regulate electrical activity and highly tunable synapses that propagate electrical and neurotransmitter-mediated signals across a network. This hardware supports a wide range of electrical dynamics that implement memory and goal-seeking behavior (and are not directly encoded by the genome, and can be modified by experience, although they have default modes of inborn activity corresponding to instincts). Other (non-neural) cell types have exactly the same ion channel, electrical synapse (gap junction), and neurotransmitter machinery. They likewise support a kind of control software, implemented in time-varying electrochemical dynamics across tissues, which underlies patterning decisions. In both cases, techniques from the field of “neural decoding” can be used to extract embedded semantics (cognitive content in the case of the brain, anatomical pre patterns in the case of tissues) from bioelectrical state readings. The computational analogy is not meant to suggest that tissues (or the brain) operate specifically via the Von Neumann architecture used by today’s computers. Interestingly,
the central nervous systems (CNS) provides important input into pattern regulation. When a nerve cord is cut and deviated to the side wall of worms
(B), the direction of the nerve end specifies whether a tail or head anatomical structure is produced. In tadpole tail regeneration
(C), laser-induced damage created within the spinal cord produces distinct changes to the shape of the regenerating tail depending on the number and location of the pinpoint holes.
Indeed, the contribution of neural signals to patterning, seen in older work on neural determination of head-tail polarity in regeneration (Fig. 2B), as well as in more recent data on the spinal cord’s role in guiding tail regeneration (Fig. 2C) and on the brain’s role in muscle and peripheral nerve patterning, helps blur the boundary between electrical activity in the brain and patterning processes in the body.
A basic introduction to developmental bioelectricityBioelectric circuits consist of ion channel proteins, which passively segregate charges across the membrane, and ion pumps, which use energy to transport ions against concentration gradients. Ion translocators in the plasma membrane set the resting potential of each cell (measured in millivolts mV). In addition to local voltage potentials, bioelectric events can be propagated across long distances by:
ephaptic field effects,
transepithelial electric fields,
tunneling nanotubes,
transfer of ion channels via exosomes,
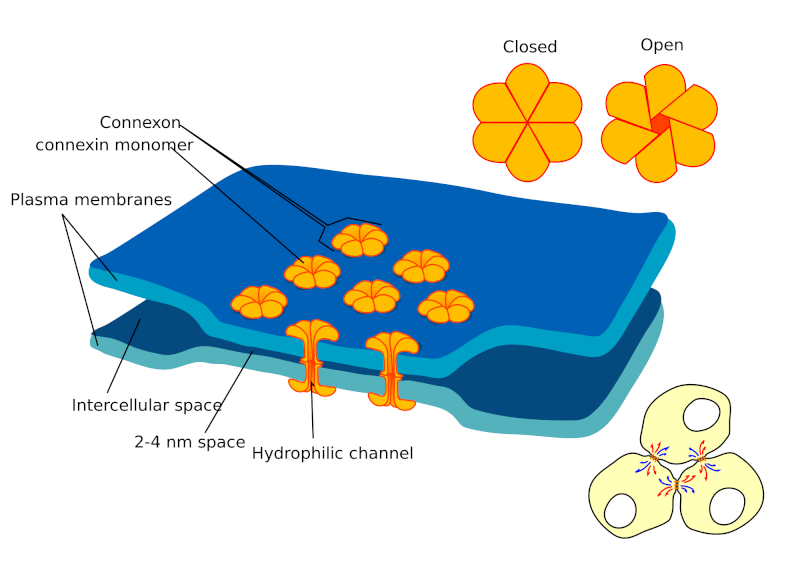
gap junctional connectivity implemented by Connexin and Innexin proteins.
Whereas ion channels and pumps exchange ions with the outside milieu, gap junctions (GJs) are channels that enable direct cell-to-cell transfer of electrical signals (and other small molecules), from the cytoplasm of one cell to its neighbor. Cells connect to each other via such electrical synapses, which facilitate the formation of bioelectrical networks that help shape the distribution of resting potential levels (Vmem) within large groups of cells and form isopotential cell fields in vivo demarcated by GJ isolation zones. “Vmem gradients” are defined as patterned spatial differences among the Vmem values of cells across anatomical distances (Fig. A below).
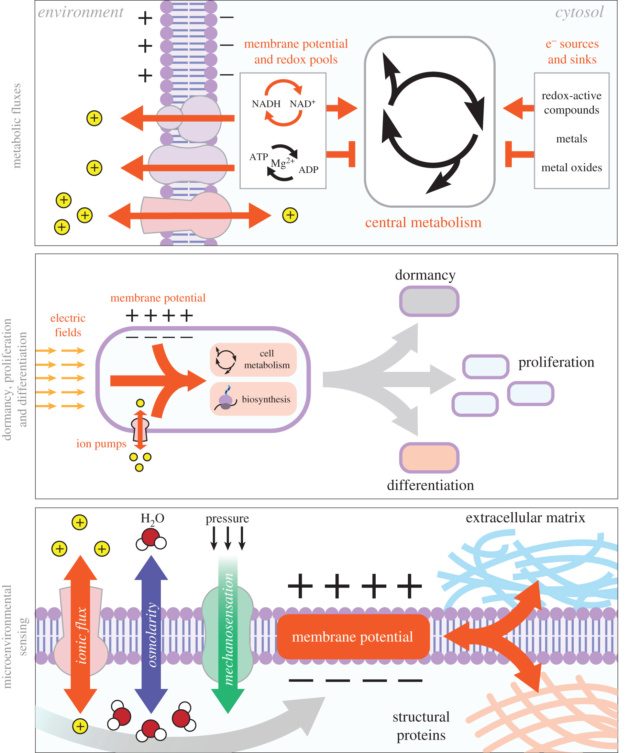
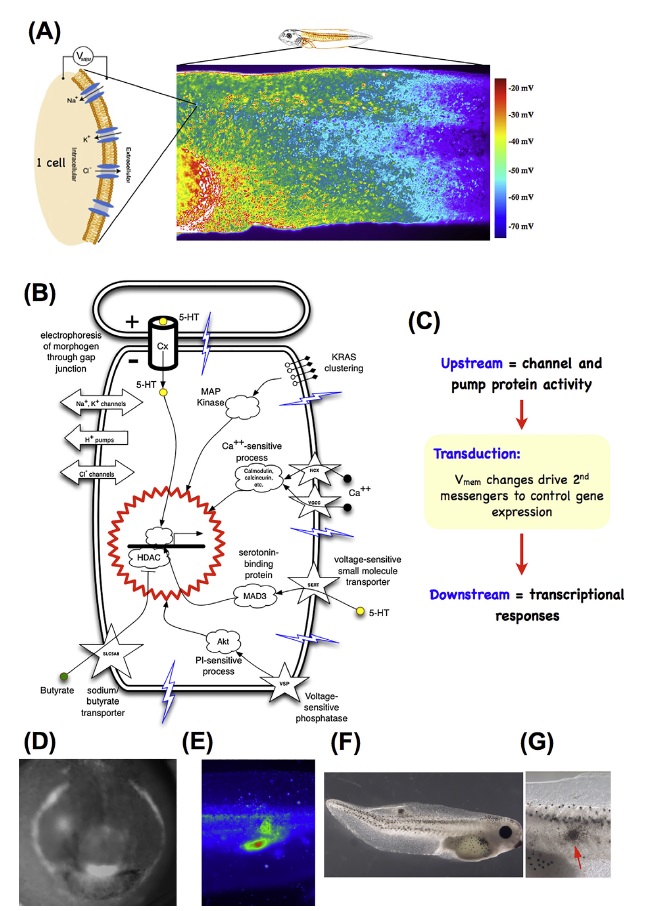 Developmental bioelectricity.(A)
Developmental bioelectricity.(A) Individual cells express ion channels and pumps in their membrane, which establish cell resting potentials. Voltage-sensitive fluorescent dyes can be used to view the Spatio-temporal patterns of these potentials in vivo, such as the flank of a tadpole seen here.
(B) Changes in voltage are transduced into second messenger cascades and downstream transcriptional responses by a variety of mechanisms including voltage-gated calcium channels, voltage-powered transporters of serotonin and butyrate, voltage-sensitive phosphatases, and electrophoresis through gap junctions.
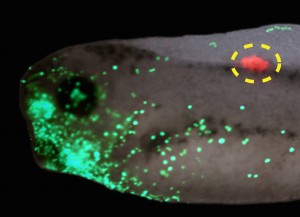
(C) Thus, the activity of ion channels and pumps are transduced into changes of gene expression(which may include other ion channels, thus forming a feedback cycle). Spatial patterns of voltage and their signaling consequences serve as pre patterns for normal morphogenesis, such as the pre patterns of the tadpole face, or disease states such as tumors induced in tadpoles by expression of human oncogenes, detected by their bioelectric disruption
(E) before they become morphologically obvious (F, close-up in G). Panel E-G
“Bioelectrical signals” are defined as a temporal change in such a pattern, which can trigger downstream patterning cascades. Cells respond to their own electrical states as well as those of their neighbors, via a range of transduction mechanisms (Fig. 3B,C;) that convert bioelectrical properties into transcriptional responses via second-messenger cascades. These include the familiar calcium pathway, and the movement of neurotransmitter molecules under voltage-gated or electrophoretic forces, precisely the same strategy used in the central nervous system. One example is the electrophoretically-controlled movement of serotonin across the early left-right axis of the developing frog embryo − a second-messenger signal that triggers asymmetric gene expression by differentially binding an intracellular receptor and chromatin modification machinery. Another example is the bioelectrically-powered movement of butyrate in and out of cells, which targets the same chromatin modifier (histone deacetylase) to regulate tumorigenesis.
What bioelectric signals do: instructive influence over morphogenesisBioelectric properties control cell behaviors, often acting as a switch between the organized collective behavior of a patterned tissue and tumorigenesis. For example, transmembrane voltage levels control the proliferation of a wide range of cell types, and Vmem regulates differentiation in a range of stem/progenitor and IPS cells. Proliferation and cell shape are known to be regulated by voltage properties, which encode important parameters at the level of individual cells. However, rich encoding properties for bioelectricity are revealed when network (multicellular)-level roles are examined.
Classic investigations of bioelectricity utilized applied electric fields via electrodes,which showed that numerous cell types read electric field lines to decode positional information and direction information for morphogenesis and migration respectively. New techniques include voltage-sensitive fluorescent dyes to read out Vmem information in vivo, and the misexpression of ligand- or light-gated ion channels and pumps to write desired voltage changes into tissues in a spatially- and temporally-controlled manner. Computational tools that provide physiology-level modeling platforms for understanding bioelectric dynamics have also been important and quantitative theory for inferring information-processing aspects of bioelectric state change. All of these developments have proceeded in parallel with similar efforts in neural decoding and inception of false memories via bioelectric editing in the field of neuroscience. The information content of bioelectrical cell states is thus an active area of investigation. One property likely mediated by these gradients is that of positional information, as long-range bioelectrical gradients are an ideal modality for coordinating spatial configuration with functional decision-making at the cell level. Future work will determine how electrically-mediated positional signaling integrates with the molecular-genetic systems of positional memory, such as the HOX code in the skin and muscle. Importantly, however, recent work has begun mechanistically linking bioelectric activity in cells with canonical pathways such as BMP.
Using a range of new strategies for functionally investigating the importance of bioelectric states, recent work has revealed that voltage gradients instruct much more than cell-level behaviors. Specific bioelectric pre patterns (Fig.D) have been found inthe face and brain, which appear to encode locations of specific anatomical features. When these pre patterns are artificially altered(Fig. E-G), predictable changes in downstream gene expression and anatomy result. Moreover, alterations of bioelectrical pattern(by introducing new ion channels, or using drugs to target endogenous channels) can induce regeneration of entire appendages in tadpoles, change the size of body structures in planaria and zebrafish,and induce reversal or randomization of the whole body axes, regulating left-right asymmetry in chick and frog and anterior-posterior anatomical polarity in planarian regeneration.All of this work clearly demonstrates that bioelectricity is an important input into pattern regulation, operating in embryogenesis, regeneration, and cancer suppression. But does it implement a code? Or is it just another set of mechanisms, like the biochemical gradients and mechanical forces that also participate in pattern regulation? Since these codes must be physically embodied, we must next decide what features constitute a code, and demarcate a set of events as best described as a coding/decoding system as opposed to apurely mechanical mechanism. A thorough philosophical discussion of principled criteria for treating systems from mechanical vs. information-processing perspectives has been presented previously.
Cracking the bioelectric codeWhy is bioelectric signaling an example of a code? We use several key features to define the existence of a “code”, in distinction to a purely mechanistic set of consecutive physical states: when does some property “encode” information, rather than merely causing downstream events? This is a complex philosophical issue related to an extensive literature on causation and semiotics,and it is likely that the distinction is a continuum without a sharp definitive distinction. Here we attempt to only give a few practical guidelines, to illustrate what is meant by a code in the context of biological patterning. The first feature focuses on the fact that a signal’s encoded meaning is not inherently tied to the physical property of said signal, but rather it is arbitrary and defined by (evolutionary) convention −amply illustrated by the extensive use of symbolic codes in human technology.
My comment: The author claims that a signal’s encoded meaning is not inherently tied to the physical property of said signal, but rather it is arbitrary and defined by (evolutionary) convention −amply illustrated by the extensive use of symbolic codes in human technology. This is blatantly false. Conventions are set by minds, by intelligence, something that evolution inherently lacks. This sets a significant paradigm shift - from evolution to intelligent setup. A signal encodes some state if its presence triggers that state to occur because of how the system interprets that signal, not because it’s a physical event that forces the outcome.
My comment: Interpretation depends on the pre-establishment of the meaning of the signal, and the assignment of meaning is always something done by intelligence. An electric field may force the movement of a charged particle − this is not an example of a code; a pattern of electric events may be interpreted by cells and cue them to differentiate − this is a code in the sense that those electrical events do not in themselves require any link to differentiation, and
evolution intelligence could have wired the system so that those same events instead signaled the need for programmed cell death. As another example, the HOX code is a code because a combination of HOX transcripts causes specific structures (e.g., wings vs. legs) to form in an organism, but none of those proteins actually have any wing- or leg-like properties or activities − it is the interpretation of this arbitrary signal by cell groups that give it meaning and a functional context. Thus, the decoding process is key. For example, a heat pulse that denatures proteins and induces cell death is not an encoded signal because its effects are physical and direct, not requiring an evolved system to interpret it.
The bio-electric code is a code in this sense because the voltage properties do not themselves imply any particular kind of patterned structure− it is the act of decoding of voltage-mediated signals by cell groups that interprets the code in constructing and remodeling anatomy.A closely-related property of codes is that the physical implementation is not as important as the information content. This is also true of the bioelectric code because it has been repeatedly found that cells can respond not to individual channel proteins or individual ion species, but to resting potential − voltage, which is an aggregate property and
can be encoded by a range of chloride, sodium, and potassium concentrations.My comment: So it is the concentration gradient that provides the relevant signals, that trigger a certain outcome. There are exceptions to this(e.g., ion channels that operate as binding proteins for other partners, and calcium, which functions in extremely small quantities a unique chemical signal rather than setting Vmem, but over-all the same outcome can be achieved by triggering appropriate Vmem states regardless of which ion translocator or ion is used. For example, tail regeneration in tadpoles can be rescued after the shut-down of the native V-ATPase proton pump by misexpression of a completely different pump from yeast that has no sequence or structural homology. Likewise, eye patterning can be induced by a range of channels that set Vmem to a specific level, while metastatic behavior in normal melanocytes can be induced or rescued using chloride, sodium, or potassium as long as the appropriate Vmem signal is provided. The bioelectric code signals by virtue of its physiological state, with no1:1 correspondence to any specific gene product.
Aside from philosophical debates on the precise definition of a“code”, the value of a metaphor lies in its explanatory power − its ability to drive new research, uncover new phenomena, and enable capabilities that competing paradigms did not facilitate. One example of a conceptual gap left by today’s paradigm concerns the most important aspect of regeneration: the stop condition. How do regenerative processes know when to stop − when the correct target morphology has been achieved? This question can be put into sharp focus with the following thought experiment (Fig. below).
 How to determine what pattern cells will build to?
How to determine what pattern cells will build to?This schematic describes a thought experiment to focus attention on the stop condition for regeneration: how do cells decide what pattern constitutes ‘correct, finished’ repair so that they can stop growth and remodeling?
(A) Different species of planaria have (and regenerate) different shapes of heads, for example, round or flat.
(B) If half of the stem cells (neoblasts) of one species are destroyed (by irradiation), and some neoblasts from another species are transplanted
(C), we can amputate the resulting worm
(D) and ask: what head shape will it regenerate? Perhaps it will be an in-between (averaged) shape, or perhaps one of the sets of neoblasts is dominant, or perhaps the head will undergo continuous and unceasing deformation as neither set of neoblasts is ever satisfied with the current shape of the head. It is important to note that none of the excellent molecular-genetic work in this field has given rise to a model that can make a prediction (or constrain) the outcome of this kind of question. This thought experiment illustrates the fact that questions of control theory, representation (encoding), and algorithmic control over regeneration have been so far largely left out of mechanistic work in pattern control.
Suppose neoblasts (adult stem cells) from two planaria with distinct head shapes are combined into one body, and that body is decapitated. What head shape will result? Would it be an average between the two, a dominant shape, or a continuously metamorphosing head (as neither population of neoblasts is ever entirely happy with the current head shape)? The striking fact is that, despite numerous high-resolution analyses of stem cell function and gene-regulatory circuits in planaria, there are no models in this field that can make a prediction on this very basic question. The understanding of regeneration and the link of known facts to the key question of how it knows what to build and when to stop exhibits a profound conceptual gap not filled by existing models, coding, or otherwise. The hypothesis that bioelectric signaling is truly implementing a code makes a number of unique predictions. What has been the benefit of treating bioelectrics as a computational layer?
Over the last 20 years, we have used concepts from a computer and cognitive science to model bioelectrics as an endogenous computational medium that processes information that must be encoded and decoded by cell groups. By suggesting the first applications of a neurotransmitter, ion channel, and electrical synapse-modifying methods to pattern regulation, this perspective has given rise to a variety of unique new findings.
Testing the predictions of a code-based view of pattern regulationThe first prediction was that bioelectric properties would encode patterns at levels above the cellular level − large-scale features of anatomy that cannot be defined at the single-cell state(Fig. below).
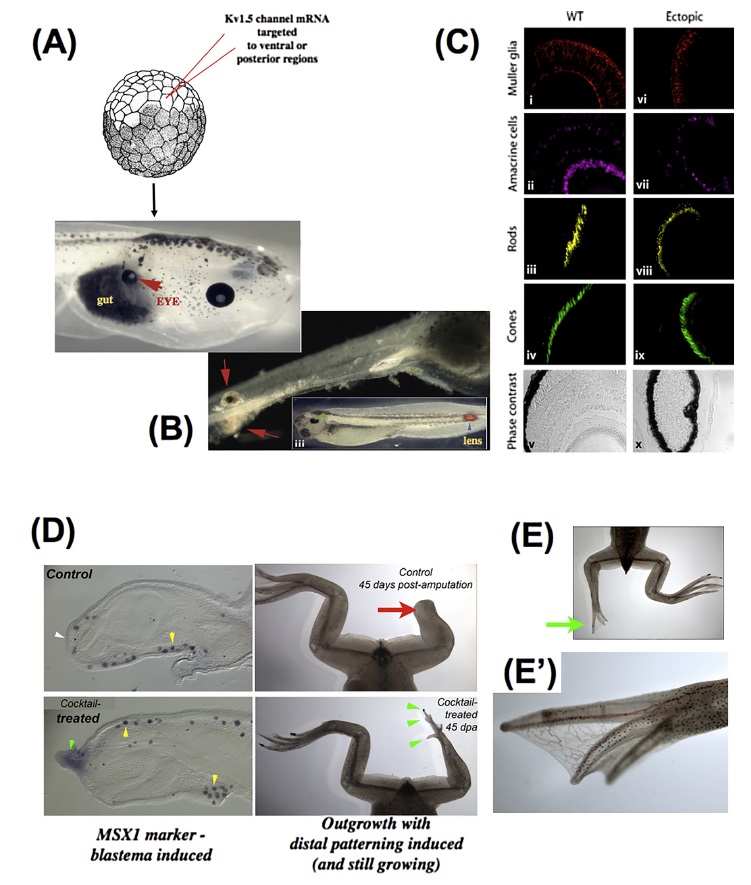 Bioelectric modification of large-scale pattern.
Bioelectric modification of large-scale pattern.Counter to early predictions that voltage outside the nervous system was a housekeeping parameter, it has been observed that specific alteration of resting potential patterns in vivo by misexpression of new ion channels can give rise to coherent, modular changes in the anatomical arrangement. In Xenopus laevis, when specific ion channel mRNAis injected into embryonic blastomeres, regions of the animal − even those outside the anterior neural field − can be induced to form a complete eye. Eye structures can be induced inside the gut (A), tail, or spinal cord (B); in some cases, these eyes possess all the correct tissue layers of a normal eye (C), revealing master-level regulator control of organ formation (triggered by a simple manipulation, not micromanagement of individual cell fates and positions). These data also reveal the ability of bioelectric signals to overcome traditional limits on tissue competence (as, for example, the master eye gene Pax6 cannot produce eyes outside the head, and it was thought that gut endoderm was not competent to form eyes). (D) Drug cocktails using blockers and activators of endogenous channels can be used to trigger a regenerative response; shown here is a monensin ionophore cocktail inducing the expression of the MSX1 gene and subsequent regeneration of the leg (including distal elements such as toes and toenails,E, close-up in E’) in a post-metamorphic frog that normally does not regenerate legs.
Experiments designed to target regions of living frog embryos with specific ion channels and thereby alter local voltage maps in a whole area during development revealed that this strategy can indeed be used to induce whole eye formation in the gut or other regions outside the head. Thus, it is possible to rewrite the morphogenetic fate of not just single cells, but tissues in a modular fashion − specifying the message “build an eye here” without needing to directly micromanage each cell’s differentiation and placement into the incredibly complex structure of a whole eye. A similar phenomenon was observed during the reprogram-ming of a planarian blastema from tail to a complete head, and in the induction of complete tail regeneration. In each of these cases, the information content provided by the exogenous channel was very small − certainly not enough to directly specify the structure of the newly-formed organ. This is one sure sign of code, in which a simple message can trigger far more complex responses after it is decoded by the recipient. This is a clear example in which such questions bear on practical issues of biomedicine: this type of “subroutine call” effect, where a simple master trigger causes a complex, self-limiting morphogenetic response, may allow regenerative medicine to provide repair of organs long before we understand everything needed to directly build a complex structure from scratch. It should be noted also that in the case of the eye, molecular-genetic master regulators, like Pax6, cannot initiate eye formation outside the anterior neural field. Thus, the bioelectric data not only reveal extensions to overly-limiting views of tissue competence that emerged from biochemical studies but enable a novel capability not previously possible to achieve.A second prediction was that the bioelectric code should allow a rich layer of control that can override genome-default outcomes. One example of this is recent work showing that contrary to the view of cancer as exclusively an irrevocable phenotype caused by clonal expansion of genetically mutated founder cells, metastatic disease can be triggered by disruption of bioelectric signaling while oncogene-induced tumors can be suppressed by enforcing appropriate bioelectric despite the fact that the oncogene is still strongly expressed.
Similarly, it was shown that brain defects induced by a mutated Notch gene can be rescued, resulting in normal brain structure, gene expression, and behavior (IQ), by artificially inducing brain-specific bioelectric pre-patterning during Xenopus development. All of these examples illustrate the dissociation of genetic state from the outcome (predictions based on transcriptomic, proteomic, or genomic analyses of each of those cases would have been incorrect), and highlight the essential nature of bioelectrically-encoded information in health and disease. A third prediction is that it should be possible to coax significantly different anatomies from the same wild-type genome,using simple perturbations that result in coherent changes in large-scale structure without requiring extensive tweaking of underlying mechanisms (Fig. A,B below).
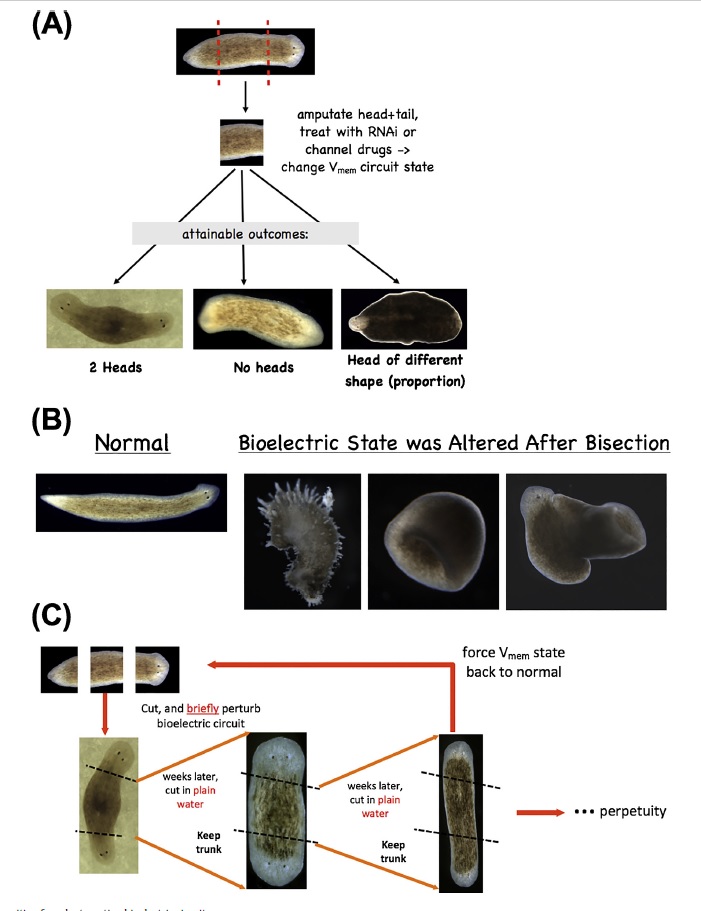 Re-writing form by targeting bioelectric circuits.(A)
Re-writing form by targeting bioelectric circuits.(A) Altering the electrical properties of tissue (by targeting ion channels, pumps, and gap junctional connectivity) in planarian fragments can be used to alter the head-tail polarity (producing double- or no-headed worms) or change the size of head structures. Moreover, such manipulations
(B) can give rise to drastically altered forms that even depart from the normal flat anatomy of planaria, all with a wild-type genomic sequence. Most importantly
(C), such changes can be permanent:double-headed planaria produced in this way continue to regenerate as double-headed when the ectopic heads are amputated, in plain water, revealing that brief (48 h)targeting of the bioelectric network to induce a different circuit state (with depolarized regions at both ends) permanently re-species the target morphology to which each piece of this worm would regenerate upon damage. The worms can be set back to normal by treatment with pump-blocking reagents that restore the normal bioelectric encoded pattern. Panels in (A) are Reproduced with permission from and. Panels in B are reproduced with permission from.
A recent example of this is the observation that temporary reduction of the bio-electric connectivity in planarian tissues during regeneration could induce a piece of
Girardia dorotocephala planarian to regenerate heads appropriate to two other species of planaria. Fragments of Girardia flatworms, treated with a gap junction uncoupler, regenerated head shapes, brain morphology, and stem cell distributions appropriate to two other extant species of planaria and completely different from their genome-default morphologies. Despite a wild-type genomic sequence, bodies can produce anatomical structures quantitatively similar to those of species 150 million years distant. The evolutionary prevalence of morphological change via changes in the bioelectric layer remains to be studied. However, the fact that other species’-specific anatomies can emerge from the same genome is likely to be an important part of understanding the relationship between genotype and anatomical phenotype during evolution for the following reason. Consider planaria; their most frequent mode of reproduction is fission followed by regeneration; thus, they escape Weissmann’s Barrier −without an obligate sperm and egg stage, somatic mutations propagate into the next generation (as long as they are not lethal to the cell). Planaria have been subject to somatic mutations and yet their morphology is robust − they regenerate correct planarian anatomy every single time, without developing cancer or aging. Moreover, planaria are the only model system in which no genetic patterning mutant lines are available. Every other model (C. elegans, zebrafish, chick, mouse, frog, etc.) offers genetic mutants with altered morphology. With only one exception(produced by a bioelectric perturbation described below), flatworm lines are always normal, perfect planaria.
How is it possible that with a highly variable genome, as befits a system in which somatic mutations are heritable, planarian regenerative and developmental processes exhibit the highest-fidelity of any other model species?
The inability of any existing data or models to answer this question (or indeed, to address the more general question of how much mutagenesis we would expect a genome to experience before anatomical change results), highlights how much remains to be learned about morphogenetic codes beyond the genetic code. All of the above examples reveal the importance of information encoded in physiological networks as a mediator of the morphogenetic code between the genome and the anatomy. Bio-electric signaling is a new type of epigenetic process, with dynamics quite distinct from that of chromatin-modifying effects as it is fundamentally distributed (multi-cellular scale) and encodes patterning, not only metabolic and differentiation state information. Finally, the most fundamental aspect of the bioelectric code model is the implication that it should be possible to edit the encoded target state to which cells build, as an alternative to revising the local rules followed by each cell. If target morphology is indeed encoded as a kind of memory for anatomical patterning, then it should be possible to re-write it. A key property of memory is its lability. Can patterns be permanently altered without genomic editing? A set of studies over the last few years has established a model of trophic memory more tractable than deer antlers and crab claws: the planarian (Fig. C above). Altering the bio-electric network of regenerating planaria, using a pharmacological reagent that leaves tissues within 24 h, produces animals that are indistinguishable from normal by anatomy, histology, key markergene expression, or stem cell distribution; however, if amputatedin plain water, a proportion of these animals give rise to double-headed worms. This stochastic state, in which the pattern to which they will regenerate has been de-coupled from their current pattern (a unique outcome in regeneration), persists indefinitely − itis stable, with no more perturbation. The only thing different about such “cryptic” animals is that their endogenous bioelectric states differ from true wild-type worms − a fact that can be decoded(readout) by human observers using voltage-reporting dyes, and by cells themselves during regeneration. Moreover, double-head worms produced by this process remain double-headed in per-petuity, despite a normal genomic sequence, revealing that target morphology can be re-written by bioelectric circuit change. Every piece of that worm acquires a different (i.e.,double-headed) encoded goal state to which it will regenerate, and this process is stable to the worm’s normal mode of reproduction(fission), revealing a novel epigenetic pathway that may be important for evolutionary plasticity. Most crucially, these manipulations reveal the existence of an encoded pattern memory, because they're-write it, altering what the tissue will do in the future (latency −a key aspect of the definition of memory).
Major knowledge gapsThe idea of electrical properties of the tissue being important, even instructive, aspects of patterning has been around for a long time; prescient classical workers like H. S. Burr long ago suspected that bioelectric pre patterns encoded anatomical states. However, modern formulations of the bioelectric code concept that cohere with the advances in information sciences, computational neuroscience, and molecular evolutionary genetics comprise an emerging field. Emerging tools are coming on-line to begin to interrogate the encoding process and structure, and the mechanisms by which this code is read and could be re-written.
One major knowledge gap is not knowing precisely what is being encoded. Models that appear appropriate to specific cases include: individual cell states (cancer, mitotic control), paint-by-number direct pre patterns for gene expression domains (craniofacial patterning), positional or size information (neural tube, tail size), or selection of discrete patterning subroutines at the organ (limb, tail)and axial polarity (left-right and head-tail decisions). It is essential to extend the existing tools to enable the writing of arbitrary bio-electric state to determine which aspects of pattern control can be written and whether the encoding of these messages are via spatial patterns of voltage, temporal fluctuations, or both. The ability to read a bioelectric pattern and predict the resulting anatomy, and conversely, to induce a bioelectric pattern that will drive morphogenesis of the desired form, is the long-term goal of the research program focused on cracking the bioelectric code. Another key area of current investigation is the ontogenic origin of the code: where do bioelectric pre patterns come from? In regeneration, and in clonal reproduction (e.g., in planarian reproductive fission), the patterns can be driven by remaining tissue − they serve as pattern memories that instruct new growth, as recently shown in flatworms. What about organisms that develop from a single fertilized egg cell?
In this case, there are two (not mutually exclusive) ways for bioelectric patterns to arise. One is via self-organization and symmetry breaking driven by amplification and feedback loops in the electric circuit. This has been well-studied for biochemical signals, and likewise occurs in electric networks (neural and non-neural). The implication is that while every multi-cellular electric circuit has a default bioelectric dynamic, this dynamic can be changed by experience (environmental or experimental perturbation) and thus diverge from the genome-default. This means that bioelectric circuits are analogous to Turing-type self-organization in the sense that they emerge from symmetry-breaking generic processes, but have additional capabilities because tunable synapses (voltage-gated channels and gap junctions) enable the stable patterns to change based on its physiological history. A second possible origin of macroscopic bioelectric patterns in development should be mentioned, although it is at present purely hypothetical. It is possible that at least some aspects of large-scale organismic bioelectric pre patterns are projected upon(represented in) subcellular domains on the surface of the egg cell. As the cells proliferate during embryonic cleavage stages, these domains would be partitioned into differential bioelectric proper-ties among distinct cells (and of course become modified beyond simple expansion by the complex feedback loops in electrically-interacting cells). It is known that cells bear many different domains on their surface, as small as a few microns in size, each of which can support distinct Vmem even though adjacent. While the functional relevance of these domains is completely unknown, it is conceivable that even the bioelectric distributions on the surface of a single cell encode important information. If true, such a phenomenon on the surface of an egg cell provides a potential mechanism for passing complex patterns across generations without the need for the pattern encoding to start de novo in each generation . Evidence for such conjecture can be sought for in future work using optogenetics and lipid raft targeting mechanisms in large eggs such as Xenopus, although it has already begun to be addressed as a computational medium in neuroscience.
Unification: neural vs. non-neural bioelectric codesHow related are developmental bioelectric codes to the more familiar electrical signaling that encodes information in the nervous system? The mechanisms that implement the bioelectric code are not new, exotic inventions. These are ancient, primitive, highly-conserved properties that had touse ion flux to drive adaptive behavior, patterning, and metabolism all in one cell. It is, therefore, no coincidence that there functions are speed-optimized in organisms with brains, using nerves to drive the behavior of muscles (and thus rapidly move the organism through 3D space) while other cell types continue to talk to each other using slower bioelectric signals (moving the body configuration through morphospace. We recently tested another prediction of this view at the level of molecular biology, by asking how much overlap there was between genes involved in patterning and those involved in memory and cognition. We extracted all the genes from the REGene database that are implicated in regeneration and subjected these to a subnetwork enrichment analysis in Pathway Studio (Elsevier) to determine what biological processes were related to this unique list. Of the more than ∼2000 transcripts collected in “model species” (e.g. chimpanzee, frog, chicken, zebrafish, fruitfly, and others) from REGene, 1309 unique transcripts were mapped successfully into Pathway Studio using gene name + Alias. The term “Regeneration” was ranked as the top bio-logical process related to the transcripts extracted from the REGenedatabase. There were a number of additional processes related to the gene set used for “regeneration”, notably learning/memory (enrichment P < 0.0001). Of the 700 transcripts related to learning/memory in the Resnet database, 177 of these overlapped with the original list from REGene, thus ∼25% of transcripts implicated in regeneration are also involved in the process of learning/memory based upon the two databases. Of the genes that mapped successfully into Pathway Studio, there was 30% overlap between the two categories.
The analysis also identified significant overlap between genes involved in learning/memory and those involved in development and cancer. Given the ancient origin of bioelectricity and the molecular conservation of mechanisms and algorithms by which the brain and body compute, how much overlap might there be between the developmental bioelectric code and the neural bio-electric code that underlies cognition?
We suggest that the bioelectric code is a linking nexus between cognitive neuroscience, developmental biology, and the field of primitive cognition. It is likely that many tissues in vivo are a kind of excitable medium, which supports morphological computation via a range of physiological signals, including bioelectric ones. If so, they are a remarkably useful proof-of-principle model for the field of unconventional computation − living tissues erase the boundary between the computational unit and the body it drives; they process information about the shape changes to make to their own structure. A computational architecture that can robustly reason about its own shape and change that shape ‘on the fly’ is the dream of today’s robotics and morphological computation communities. Indeed, recent studies have shown that brains retain information despite drastic remodeling − this is seen in the work on persistence of memory during the transition from caterpillar to butterfly (in which the brain is largely destroyed and recreated), and in recent work confirming McConnell’s early studies showing that when trained planaria are decapitated, their tails grow new heads and remember their original learning.
The inter-face of information encoding in brain and body, especially during contests such as regeneration of the CNS in which both episodic and somatic memories must play a role, reveals the lack of a sharp dividing line between neuroscience and pattern formation.Information encoding in living tissue is the fundamental umbrella under which regenerative biology, neuroscience, robotics, etc. all operate. It is likely that conceptual tools of computational neuroscience will help crack the patterning bioelectric code, for example by determining how spatial patterns can be represented as memories within a (non)neural network (Fig. below).
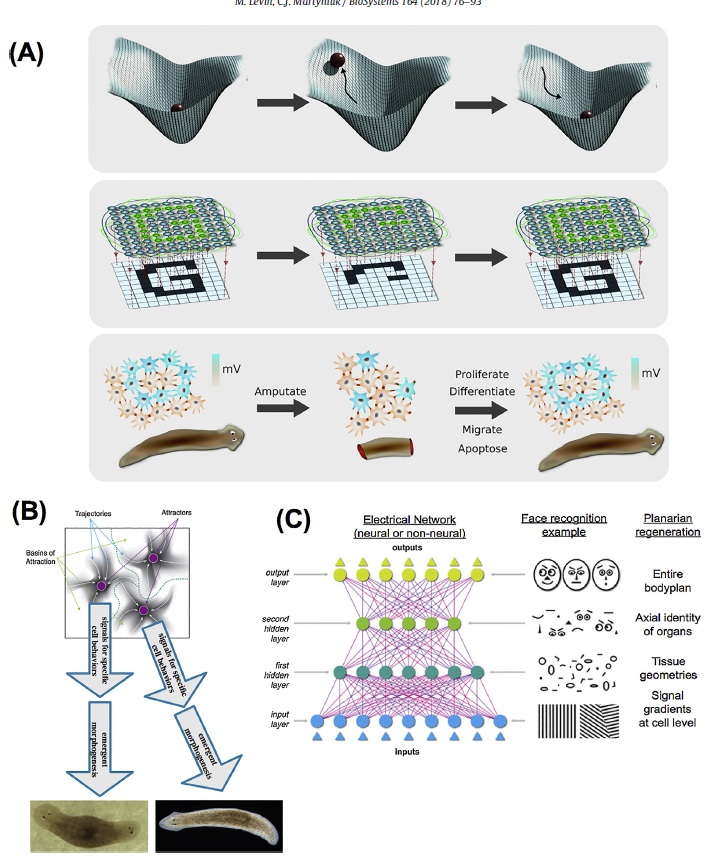 A neural network view of bioelectric circuits.(A)
A neural network view of bioelectric circuits.(A) One way to think about the regenerative (pattern-homeostatic) process is as a dynamical system in which the correct shape represents an attractor. Damage raises the energy of the system (here represented by the ball leaving the lowest-energy state), but it settles back to the correct state by a least-action mechanism minimizing error. The field of artificial neural networks (ANNs) and computational neuroscience, which have begun to explain the holographic storage of patterning information in networks, provides a number of conceptual frameworks in which to understand the function of non-neural bioelectric networks as implementing a distributed,self-correcting pattern mechanism.
(B) Specifically, in certain kinds of networks, attractors in their state space correspond to specific memories.
We propose a model in which attractor states of bioelectric circuits correspond to specific anatomical layouts by controlling cell behaviors such as proliferation and differentiation. This provides a quantitative, mechanistic approach to understand how electrical signaling encodes pattern memories. Deforming the landscape, or altering cells’ interpretation of the network’s instructions, are both ways to manipulate the outcome.
(C) Another important insight provided by the field of artificial neural networks is that of encoding“high level” items: middle layers of ANNs encode emergent features of the inputs; this provides a way to think about how cell signaling networks (in particular, electrical networks) could encode and decode information about parameters above the single-cell level (organ size, topological arrangement, etc.).
It is just as likely that progress on decoding the simpler (?) encoding of patterns in non-neural contexts will help efforts to develop neural decoding (i.e., to extract first-person cognitive content from brain scans). Integrated cases (such as storage of learned memories in the bodies of planaria and their imprinting on the nascent brain)are expected not only to enrich our understanding of the material embodiments of mind but to also drive transformative applications in-memory technology. Research in regenerative biology is already driving extensions of computer science, such as the investigations of the stability properties of artificial neural networks under topology change; this has only begun to be explored and is not only useful as a model for traumatic brain injury repair and computational psychiatry but also as an extension of the artificial neural network (ANN) paradigm and connectionist theory in general to non-neural substrates
Likewise, we cannot understand the development process until we really understand the relationship between genomes and anatomy. It is consequently of the utmost importance to understand how shape is encoded in cellular properties and learn to control that shape. Ever-finer reductive drill down into single-cell molecular events is not going to be sufficient; a complementary synthetic effort to understand the algorithms and encoded meaning (not just the mechanisms) of pattern regulation is essential.
Alongside the genetic and (chromatin) epigenetic codes operate a crucial set of controls called the bioelectric code. While the encoding of pattern outcomes to bioelectric states is only known fora small handful of examples, it is clear that profound lessons about the source and nature of the information that determines anatomical pattern remain to be learned. Manipulation of long-term, the large-scale anatomical structure can now be achieved by transiently-writing the bioelectric states of living tissue. Work in tractable animal models is now beginning to be done in human cells, and this field has many applications in human therapeutics aimed at cellular control and the creation of novel bioengineered constructs.
Bioelectricity is not just another pathway. It is uniquely suited for the control of complex outcomes (encoding) because voltage-sensitive ion channels and electrical synapses implement signaling elements that are sensitive to history (voltage-gated ion currents −in effect, transistors). Those elements are easily turned into memory circuits and logic gates, which in turn enable flexible, robust computational properties. Still, it is not just about bioelectricity; ionic circuits are only one layer of the morphogenetic code (Fig. below).
 The relationship of the bioelectric and genetic code.
The relationship of the bioelectric and genetic code.The genetic code specifies proteins via gene-regulatory networks, the activity of which results in emergent patterning events. Coupled to this (but having its own distinct dynamics and capabilities) is the bioelectric code, in which cell networks use direct, indirect, or neural-like representations of specific patterns at different scales of the organization to specify instructions for modifying anatomy. Together, these codes serve as pattern memory.
The main point is that biological pattern regulation is a combination of emergent features that fill in local features and top-down controls that make decisions about large-scale patterning. One direction for future work is to attempt to re-write the encoded goal state, instead of manipulating individual cells; to the extent that cells may be “universal constructors”, deforming their perception space landscape and letting them do what they are good at (finding stable attractors within that space), might be a most efficient path to pattern control.
Major opportunities in this field include the development of quantitative theory − conceptual models based on dynamical systems theory, least-action field-like principles, and Bayesian inference.
Integrating the fundamentals of bioelectrical dynamics with the increasing progress in the understanding of gene-regulatory networks and physical forces in patterning will be an essential next step. One important aspect however is scale-invariance: numerous aspects of pattern control, such as intercalation of positional information values, appear to work similarly in single cells as they do in multicellular organisms. Thus, it may be possible to derive fundamental laws in which patterning out-comes can utilize diverse underlying mechanisms to implement them. If indeed information processing is ancient, one of such fundamental laws may be the drive to reduce high-level measurable such as surprise. Cells may have an innate capacity to learn to anticipate their environment and act in a way that maximizes reward; this has already been shown for cultured neurons in vitro, which can learn to operate a virtual flight simulator “body”. Thus, direct training of non-neural tissues by providing positive and negative reinforcement for patterning behavior could be a way to offload the computational complexity of patterning tasks from the bioengineer onto the living system, rewarding for desired behavior and letting the system self-organize internal processes to achieve the necessary goal. This kind of generalized plasticity in service to specific outcomes closely related to a key insight that drove the development of the computer science revolution − the independence of hardware and software, and the ability to run the same software on different hardware, or obtain different behavior from the same hardware by changing the software. If bioelectric dynamics running on genome-specified ion channel complements in cells can be treated as a kind of software, the next revolution in biology could be likewise driven in part by the realization that we do not have to manipulate living systems at the level of their “machine code” (affecting specific molecules), but at the level of information − re-writing the encoded goal states and thus gaining a more top-down control overgrowth and form with myriad applications to biomedicine and robust technology
https://sci-hub.ren/https://www.sciencedirect.com/science/article/abs/pii/S0303264717302848