10. Protoporphyrin IX Mg-chelatase(1) glutamyl-tRNA synthetase; (2) glutamyl-tRNA reductase; (3) glutamate 1-semialdehyde aminotransferase;(4) porphobilinogen synthase; (5) hydroxymethylbilane synthase; (6) uroporphyrinogen III synthase;(7) uroporphyrinogen III decarboxylase; (8 ) coproporphyrinogen III oxidative decarboxylase; (9) protoporphyrinogen IX oxidase; (10) protoporphyrin IX Mg-chelatase; (11) S-adenosyl-L-methionine:Mg-protoporphyrin IX methyltransferase;(12)–(14) Mg-protoporphyrin IX monomethyl ester oxidative cyclase; (15) divinyl (proto)chlorophyllide 4-vinyl reductase; (16) light-dependent NADPH:protochlorophyllide oxidoreductase or light-independent protochlorophyllidereductase; (17) chlorophyll synthase.
Magnesium-chelatase is a
three-component enzyme that catalyses the insertion of Mg2+ into protoporphyrin IX. This is the first unique step in the synthesis of chlorophyll and bacteriochlorophyll.[1][2] As a result, it is thought that Mg-chelatase has an important role in channeling intermediates into the (bacterio)chlorophyll branch 1
The reaction: 5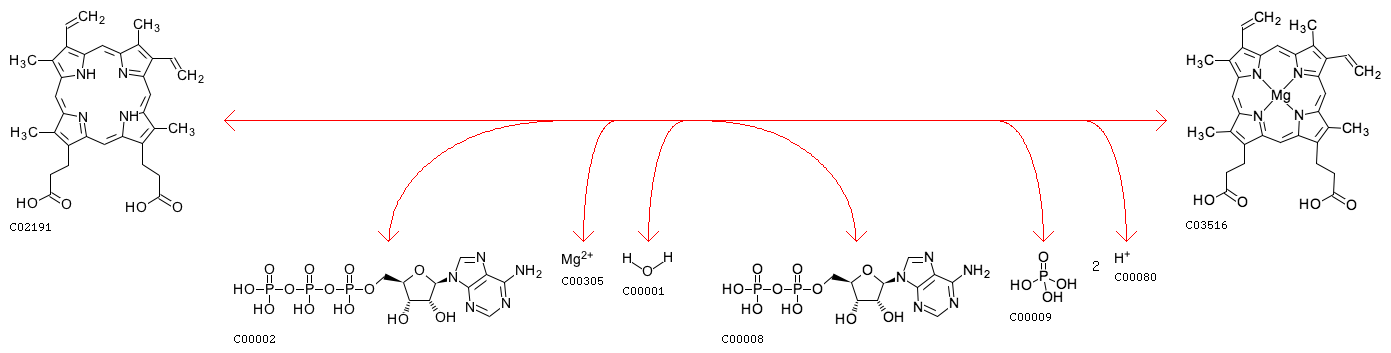 Amino Acid sequence
Amino Acid sequence 863 aa
Nucleotide sequence: 2589 nt
https://www.genome.jp/entry/ajm:119045164
Robert D. Willows The Mg branch of chlorophyll synthesis: Biosynthesis of chlorophyll a from protoporphyrin IX 2019 1The protein subunits are generally known as
I, D and
H with the prefix “Bch” and “bch” to the proteins and encoding genes, respectively, from bacteriochlorophyll-synthesizing organisms and “Chl” and “chl” to the proteins and encoding genes, respectively, from chlorophyll-synthesizing organisms
All subunits are highly conserved among bacteria and plants, with about 60 % of the amino acid positions invariant in all known sequences of BchI, 30 % in BchD and 40 % in BchH. 2
The
I protein sequences belong to a unique protein family (pfam), has
41% sequence homology across the family. ( homology is similarity due to shared ancestry between a pair of structures or genes in different taxa.
a )
The 2.1A˚ X-ray structure revealed that BchI belonged to the AAA+ class of proteins with a
novel inverted domain structure linked by a long kinked alpha helix.
In contrast, the
D proteins are
quite divergent in their sequences and they do not belong to a single pfam. The D proteins have an AAA+ (ATPases-associated with diverse cellular activities’)-N-terminal domain, similar to the I protein, which is linked to an integrin-I/von Willibrand C-terminal domain by a polyproline sequence containing 8–10 prolines followed by a low complexity highly charged domain.
The H protein sequences belong to a
unique pfam, consisting of magnesium chelatase H and cobalt chelatase CobN subunits.
The four substrates of this enzyme are ATP, protoporphyrin IX, Mg2+, and H2O; its four products are
ADP, phosphate, Mg-protoporphyrin IX, and H+. This enzyme belongs to the family of
ligases, specifically those forming nitrogen-D-metal bonds in coordination complexes.
The first step in the synthesis of chlorophyll from PPIX is the insertion of magnesium into the tetrapyrrole macrocycle to make magnesium proto porphyrin IX (MgPPIX). Magnesium chelatase
requires a minimum of three protein subunits for activity and
requires ATP hydrolysis and the
substrates Mg2+ and PPIX. An accessory protein,
GUN4, is also important for regulating magnesium chelatase activity in oxygenic photosynthetic organisms.
 Dominique Pontier Knock-out of the Magnesium Protoporphyrin IX Methyltransferase Gene in Arabidopsis JANUARY 2007 3
Dominique Pontier Knock-out of the Magnesium Protoporphyrin IX Methyltransferase Gene in Arabidopsis JANUARY 2007 3Protoporphyrin IX is the last common intermediate between the heme and chlorophyll biosynthesis pathways. The addition of magnesium directs this molecule toward chlorophyll biosynthesis. The first step downstream from the branchpoint is catalyzed by the magnesium chelatase and is a
highly regulated process. My comment: According to Wiki 4 Regulation is the
management of complex systems according to a set of rules and trends. In systems theory, these types of rules exist in various fields of biology and society, but the term has slightly different meanings according to context. For example: In biology, gene regulation and metabolic regulation allow living organisms to adapt to their environment and maintain homeostasis; Was regulation of the process not something necessary right from the start ? That means, it would have had to evolve in a coordinated sense together with the enzyme activity, and afterward, somehow, brought together to work in a functional relationship? But in order to coordinate something, foresight is necessary. Something that mindless evolutionary processes lack.
The corresponding product, magnesium protoporphyrin IX, has been proposed to play an important role as a signaling molecule implicated in plastid-to-nucleus communication. To get more information on the chlorophyll biosynthesis pathway and on magnesium protoporphyrin IX derivative functions, we have identified an magnesium protoporphyrin IX methyltransferase (CHLM) knock-out mutant in Arabidopsis in which the mutation induces a blockage downstream from magnesium protoporphyrin IX and an accumulation of this chlorophyll biosynthesis intermediate. Our results demonstrate that
the CHLM gene is essential for the formation of chlorophyll and subsequently for the formation of photosystems I and II and cytochrome b6f complexes.
Filipa L. Sousa Chlorophyll Biosynthesis Gene Evolution Indicates Photosystem Gene Duplication, Not Photosystem Merger, at the Origin of Oxygenic Photosynthesis 19 December 2012 6Mg ChelataseThe first unique intermediate of chlorophyll biosynthetic pathway, Mg-protoporphyrin IX, is generated by the insertion of Mg2+ into protoporphyrin IX. Biochemical and genetic analysis identified a Class I ATP-dependent magnesium chelatase, composed of three subunits BchH, BchI, and BchD, that catalyzes a reaction consisting of an activation and a chelation step. In the presence of both ATP and Mg2+, an AAA+ motor complex is assembled from a hexameric or heptameric BchI ring connected to a hexameric BchD ring (the activation step). Proto IX binds to the BchH catalytic subunit, and its transient interaction with the formed AAA+ motor complex leads to the insertion of Mg2+ in the tetrapyrrole macrocycle (the chelation step).
The Mg-chelatase complex has both sequence and structural homology with the Class I cobalt chelatase of the O2-dependent cobalamin biosynthetic pathway. The CobN, CobS, and CobT subunits of the trimeric cobalt chelatase are homologous with the BchH/ChlH, BchI/ChlI, and BchD/ChlD subunits of magnesium chelatase, respectively. This homology has been interpreted as reflecting ancient duplication and divergence. Although there is a broad taxonomic distribution of the CobN gene among cobalamin-dependent organisms,
a much narrower distribution is observed for the genes that compose the AAA+ motor complex (CobS and CobT).
The Catalytic Subunit BchH/ChlHChlorobaculum tepidum, similar to most members of GSB, has three paralogous genes ( Paralogous genes are genes that are related via duplication events in the last common ancestor (LCA) of the species being compared. ) for this subunit, named BchS, BchH, and BchT. Single- and double-mutant experiments showed that strains with only the BchH or the BchS gene retained sufficient Mg-chelatase activity to be viable, but its activity was maximal if the BchS gene was present together with BchH. This was also confirmed by biochemical characterization of the recombinant enzymes. However, in all mutagenesis experiments, a decrease in the production of bacteriochlorophyll c, the main photosynthetic pigment of GSB, was detected. This suggests that
the different isoforms ( protein variants) function in end-product regulation and/or substrate channeling of the Bch c intermediates.
The existence of BchH/ChlH paralogs is not unique to the GSB. All chloroflexi have at least one additional gene coding this subunit, and the same is also true for some cyanobacterial species. Isoforms of the gene also occur in some eukaryotes although, in this case, different duplication events are possibly at their origin.
The AAA+ Motor: BchI/ChlI and BchD/ChlDThe BchI/ChlI subunit is a member of the AAA+-ATPase family of proteins, which includes proteins of diverse function. With the exception of some land plants and green algae, there is a consensus in the literature that photosynthetic organisms only have one BchI/ChlI subunit. However, a study of two recombinant isoforms of BchI from Prosthecochloris vibrioformis showed that both had Mg chelatase activity in vitro, and in the recently sequenced genome of Chloroflexus aurantiacus, three genes were annotated as BchI. In our search, several isoforms belonging to both chlorobia and chloroflexi were retrieved. These enzymes are distinct from the aerobic cobalamin chelatase CobS gene and were included in the analysis.
The BchI/ChlI tree is divided into two clades, one comprising BchI/ChlI genes among all photosynthesizers and one containing the different isoforms from GSB and GNSB, which we call the GSB/GNSB clade. In the former, two major groups can be observed, one with heliobacteria, chlorobia, chloroflexi, acidobacteria, and proteobacteria (branching in that order) and a second one, where the cyanobacteria and eukaryotic sequences cluster together. The isoforms present in the GSB/GNSB clade, as in the case of G. violaceus ChlH2 sequence, have high similarity with sequences similar to hypothetical magnesium chelatases from nonphotosynthetic organisms. Although the results from P. vibrioformis indicate that both isoforms have identical activities in vitro, their sequences show higher similarity with sequences from organisms that do not synthetize bacteriochlorophyll, questioning their role in chlorophyll biosynthesis. Specifically, within delta-proteobacteria, proprionobacterales, firmicutes, and euryarchaeota, some organisms possess isoforms of the BchI/ChlI gene in addition to CobS. Other cobalamin producers possess only the BchI/ChlI gene and lack CobS. In these organisms, the BchI/ChlI genes probably substitute the missing CobNST genes. The distributions of BchI/ChlI and CobS homologs suggest a way in which pre-existing building blocks could have been recruited into the assembly of the ancestral chlorophyll and O2-dependent cobalamin pathway.
The N-terminus of BchD/ChlD gene, which is also a member of the AAA+-ATPase family of proteins, has a segment of approximately 260 amino acid residues with sequence homology to BchI/ChlI. BchD/ChlD has a proposed structural role in magnesium chelatase, functioning as a platform for the convergence of the other two subunits. The BchD/ChlD phylogenetic tree in figure 3C was rooted by two sequences of von Willebrand factor Type A, which is considered to be an ancient protein domain. The overall BchD/ChlD tree topology is very similar to the upper part of the BchH/ChlH tree with the exception of the position of C. Chloracidobacterium thermophilum. There are two distinct BchD/ChlD clades. One contains acidobacteria, the GSB/GNSB clade, and proteobacteria. In the second, heliobacteria, cyanobacteria, and eukaryotic sequences cluster. There are two eukaryotic clades, one composed of green algae isoforms that branch between heliobacteria and cyanobacteria, and a second in which at least one copy from every eukaryotic organism is represented in a sister group of the G. violaceus sequence. However, this branch is poorly supported. As in the case of BchI/ChlI, copies of this gene are also present in nonphotosynthetic organisms.
Xuemin Chen Crystal structure of the catalytic subunit of magnesium chelatase 24 August 2015 7
Tetrapyrroles, including haem and chlorophyll, play vital roles for various biological processes, such as respiration and photosynthesis, and their biosynthesis is critical for virtually all organisms. In photosynthetic organisms, magnesium chelatase (MgCh) catalyses insertion of magnesium into the centre of protoporphyrin IX, the branch-point precursor for both haem and chlorophyll, leading tetrapyrrole biosynthesis into the magnesium branch. This reaction needs a cooperated action of the three subunits of MgCh: the catalytic subunit ChlH and two AAA+ subunits, ChlI and ChlD. To date, the mechanism of MgCh awaits further elucidation due to a lack of high-resolution structures, especially for the ∼150 kDa catalytic subunit. Here we report the crystal structure of ChlH from the photosynthetic cyanobacterium Synechocystis PCC 6803, solved at 2.5 Å resolution. The active site is buried deeply inside the protein interior, and
the surrounding residues are conserved throughout evolution. This structure helps to explain the loss of function reported for the cch and gun5 mutations of the ChlH subunit, and to provide the molecular basis of substrate channelling during the magnesium-chelating process. The structure advances our understanding of the holoenzyme of MgCh, a metal chelating enzyme other than ferrochelatase.
Chlorophyll, the most abundant pigment in plants, algae and cyanobacteria, is synthesized through a multistep pathway in which protoporphyrin IX (Proto) is the branch-point precursor for both haem and chlorophyll. In contrast to the single-subunit ATP-independent ferrochelatase that catalyses insertion of a ferrous iron into the Proto ring6 , MgCh comprises three subunits, ChlH, ChlI and ChlD, and requires ATP for magnesium chelation. To complete a catalytic cycle,
the three subunits cooperate in a dynamic manner with the AAA+ subunits ChlI and ChlD assembling into a two-tiered hexameric ring, and then interacting with the substrate-binding ChlH subunit to form a transient holoenzyme complex .
 Crystal structure of the ChlH subunit. a,
Crystal structure of the ChlH subunit. a, Three surface models related by 90° rotation along the vertical axis. The head (I) domain is blue, the neck (II) domain in green, domain III in orange, the insertion (IV) domain in brown, domain V is magenta, and domain VI is purple.
b, Schematic representation of Synechocystis ChlH subunit. Regions not visible in the crystal structure include residues 140–156, 221–229, 327–330, 380–391, 415–424 and 630–640. The boundary between domains I and II that cannot be defined (residues 221–229) is grey.
c, Domains I–VI as ribbon diagrams
Joakim Lundqvist ATP-Induced Conformational Dynamics in the AAA+ Motor Unit of Magnesium Chelatase MARCH 10, 2010 8
Mg-chelatase catalyzes the first committed step of the chlorophyll biosynthetic pathway, the ATP-dependent insertion of Mg2+ into protoporphyrin IX (PPIX). Here we report the reconstruction using single-particle cryo-electron microscopy of the complex between subunits BchD and BchI of Rhodobacter capsulatus Mg-chelatase in the presence of ADP, the nonhydrolyzable ATP analog AMPPNP, and ATP at 7.5 Å, 14 Å, and 13 Å resolution, respectively. We show that
the two AAA+ modules of the subunits form a unique complex of 3 dimers related by a three-fold axis. The reconstructions demonstrate substantial differences between the conformations of the complex in the presence of ATP and ADP, and suggest that the C-terminal integrin-I domains of the BchD subunits play a central role in transmitting conformational changes of BchI to BchD. Based on these data a model for the function of magnesium chelatase is proposed.
IntroductionThe enzyme magnesium chelatase (Mg-chelatase) is active in the branch point between chlorophyll and heme biosynthesis. It catalyzes the insertion of Mg2+ into protoporphyrin IX (PPIX), which is the first committed reaction of the chlorophyll biosynthesis pathway. Mg-chelatase belongs to the class of AAA+-type chelatases. This is also true for aerobic cobaltochelatase and nickel chelatase, which are active in cobalamin (vitamin B12) and coenzyme F430 biosynthesis, respectively. Magnesium chelatase is currently the most extensively studied enzyme in this class of AAA+ chelatases. Its activity requires the presence of three subunits, BchH, BchD, and BchI, which have molecular masses of approximately 140, 70, and 40 kDa, respectively. Biochemical studies have suggested that
the enzymatic reaction proceeds in distinct steps.
Question: Had these steps not to be fully functional, and intermediates would be non-functional, and not selected by natural selection, since not confering any advantage of function, nor survival ?
In the first step, the AAA+ motor complex between subunits BchI and BchD is formed in the presence of ATP and Mg2+, while the largest subunit, BchH, binds PPIX by an unknown mechanism. Our recent work has suggested that subunit
BchD may serve as a platform for the assembly of the complex. Binding of PPIX to BchH induces a large conformational rearrangement in this subunit.
In the reaction step that follows, the BchH:PPIX complex interacts with the BchI:BchD complex, leading to insertion of Mg2+ into PPIX. During this part of the reaction, the BchH:PPIX complex is a substrate of the BchI:BchD complex. It has been estimated that around 15 ATP molecules may be required for each catalytic cycle. The BchI subunit, which contains the characteristic ATP binding Walker A and Walker B motifs (GX4GKSX6A and hhhhD(D/E), where h is any hydrophobic residue), is responsible for ATP hydrolysis. The X-ray crystallographic structure of Rhodobacter capsulatus BchI has been determined and it has been shown to belong to the AAA+ family of ATPases. The protein was later assigned to the pre-sensor II (PS-II) insert clade of the AAA+ family, which includes the MCM (minichromosome maintenance) family of helicases, the MoxR family of molecular chaperones, and the dynein/midacin family of ATP-dependent motors, the members of which are known to interact with microtubules and the nuclear pore complex. A characteristic feature of AAA+ proteins is the formation of oligomeric ring structures with the most common ring types consisting of 6 or 7 monomers. Electron microscopy (EM) and single-particle analysis has indeed shown that in the presence of ATP, R. capsulatus BchI and the corresponding subunit ChlI from Synechocystis sp. PCC6830 can form hexameric and heptameric ring structures, respectively.
Amino acid sequence analysis has demonstrated that subunit BchD, which is the second in size after BchH, has an AAA+ module at its N terminus with distinct homology to BchI. However, the Walker A and Walker B motifs, which are necessary for ATP hydrolyzing activity, are poorly conserved in this subunit. Despite this, BchD is still capable of forming oligomeric ring structures, even in the absence of ATP. Interestingly, the C-terminal part of BchD was found to contain a domain homologous to a class of proteins termed integrin I domains, which is a subgroup of a larger group of von Willebrand factor A domain proteins. These domains are usually found as part of larger complexes, and they are the principal receptors on the surfaces of animal cells, being involved in the binding of most extracellular matrix proteins. Integrin I domains are characterized by the MIDAS motif (metal ion-dependent adhesion site), which constitutes a unique Mg2+/Mn2+ binding site. The same type of domain is present in cobaltochelatase and in the MoxR family of AAA+ proteins. Mutation of residues in the MIDAS motif of R. capsulatus BchD (D385A and S387A) was found to abolish Mg-chelatase activity. The integrin I domain and the N-terminal AAA+ module of BchD are linked to each other by a proline and an acidic residue-rich region. These types of domains are often involved in protein-protein interactions. Based on this knowledge, the region was suggested to be involved in the stabilization of the BchI:BchD complex.
a https://en.wikipedia.org/wiki/Homology_(biology)
1. http://pfam.xfam.org/family/PF01078
2. https://www.sciencedirect.com/science/article/abs/pii/S002228360194834X
3. https://www.jbc.org/article/S0021-9258(20)72099-5/fulltext
4. https://en.wikipedia.org/wiki/Regulation
5. https://www.genome.jp/entry/R03877
6. https://academic.oup.com/gbe/article/5/1/200/729709
7. https://www.nature.com/articles/nplants2015125 https://sci-hub.yncjkj.com/10.1038/nplants.2015.125
8. https://www.cell.com/structure/fulltext/S0969-2126(10)00031-6?_returnURL=https%3A%2F%2Flinkinghub.elsevier.com%2Fretrieve%2Fpii%2FS0969212610000316%3Fshowall%3Dtrue








































 S-adenosyl-L-homocysteine + magnesium protoporphyrin IX 13-methyl ester
S-adenosyl-L-homocysteine + magnesium protoporphyrin IX 13-methyl ester