Fatty Acid and Phospholipid Biosynthesis in Prokaryotes
In 1956 Kennedy and Weiss published the seminal paper in the phospholipid field that formed the biological foundation for future studies in all organisms thereby establishing the “Kennedy Pathways” for phospholipid synthesis.
There is a large diversity in the structures and metabolism of membrane lipids, and not a single paradigm can be considered typical.
As we now know, phosphatidic acid (PA1, 1,2 diacyl-sn-glycerol-3-phosphate (G3P) ) is the precursor leading to the biosynthesis of the remaining phospholipids in Bacteria and Eukarya.
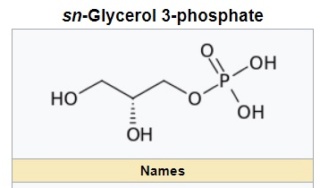
sn-Glycerol 3-phosphate is a phosphoric ester of glycerol, which is a component of glycerophospholipids. 3
Although the spectrum of phospholipid headgroup structures produced by bacteria is large, the key precursor to all of these molecules is phosphatidic acid (PtdOH). 2 Glycerol-3-phosphate derived from the glycolysis via glycerol-phosphate synthase is the universal source for the glycerol backbone of PtdOH.
My comment: This is another catch22 conundrum. Glycolysis requires a functional membrane to function. But modern membranes depend on membrane precursor phosphatidic acid to be synthesized. Another example of why cells are irreducibly complex and interdependent.
The formation of G3P from the reduction of dihydroxyacetone phosphate by the G3P synthase (GpsA) is the only de novo pathway to G3P in bacteria
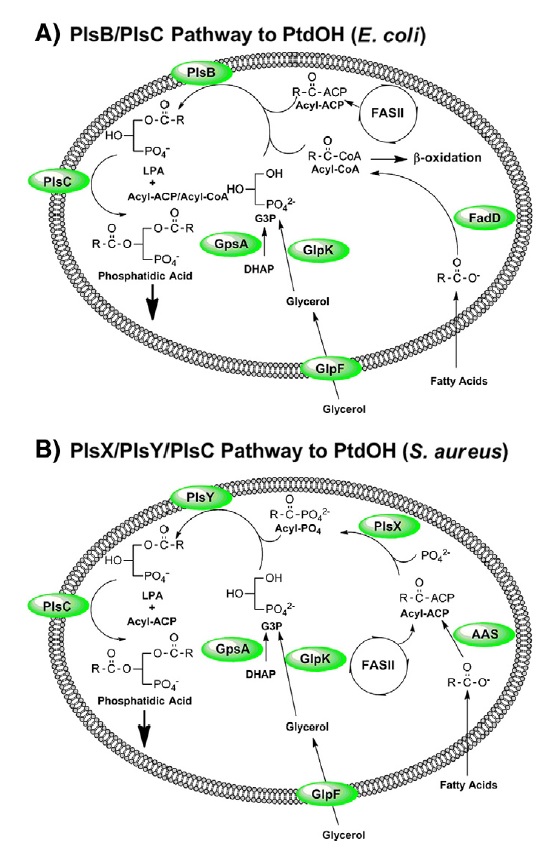
Pathways for the biosynthesis of PtdOH in bacteria.
(A) PtdOH metabolism in E. coli is representative of the bacteria that utilize the PlsB/PlsC acyltransferase pathway to PtdOH. These acyltransferases use either acyl-ACP substrates produced by type II fatty acid synthesis (FASII) or acyl-CoA thioesters generated by the activation of exogenous fatty acids by acyl-CoA synthetase (FadD). The PlsB pathway is largely confined to the γ-proteobacteria.
(B) PtdOH metabolism in S. aureus is representative of bacteria that utilize the PlsX/PlsY/PlsC acyltransferase pathway to PtdOH. Acyl-ACP generated by FASII is either used by PlsC to acylate the 2-position of LPA or is converted by PlsX to acyl-PO4 for incorporation into the 1-position by PlsY. This is the only pathway present in most Gram-positive pathogens. In both schemes, the G3P backbone is produced by GpsA, but may also be obtained from the environment by the GlpF/GlpK pathway. Many bacteria also have a transport system for G3P (not shown). Exogenous fatty acids are incorporated into phospholipid following their ligation to ACP by acyl-ACP synthetases (AAS). PlsB, PlsC, PlsY and GlpF are intrinsic membrane proteins. AAS, FadD and PlsX are soluble proteins that are thought to interact with the membrane interface.
There are two distinct families of enzymes responsible for the acylation of the 1-position of glycerol-3-phosphate. The PlsB acyltransferase was discovered in Escherichia coli, and homologs are present in many eukaryotes. This protein family primarily uses acyl–acyl carrier protein (ACP) endproducts of fatty acid synthesis as acyl donors, but may also use acyl-CoA derived from exogenous fatty acids. The second protein family, PlsY, is more widely distributed in bacteria and utilizes the unique acyl donor, acyl-phosphate, which is produced from acyl-ACP by the enzyme PlsX. The acylation of the 2-position is carried out by members of the PlsC protein family. All PlsCs use acyl-ACP as the acyl donor, although the PlsCs of the γ-proteobacteria also may use acyl-CoA. Phospholipid headgroups are precursors in the biosynthesis of other membrane-associated molecules and the diacylglycerol product of these reactions is converted to PtdOH by one of two distinct families of lipid kinases. The central importance of the de novo and recycling pathways to PtdOH in cell physiology suggest that these enzymes are suitable targets for the development of antibacterial therapeutics in Gram-positive pathogens. This article is part of a Special Issue entitled Phospholipids and Phospholipid Metabolism.
Archaea phospholipids differ in that they are composed of sn-glycerol-1-phosphate in ether linkage at the 2- and 3-positions to long-chain poly isoprenoids. 1
Synthesis of phosphatidic acid
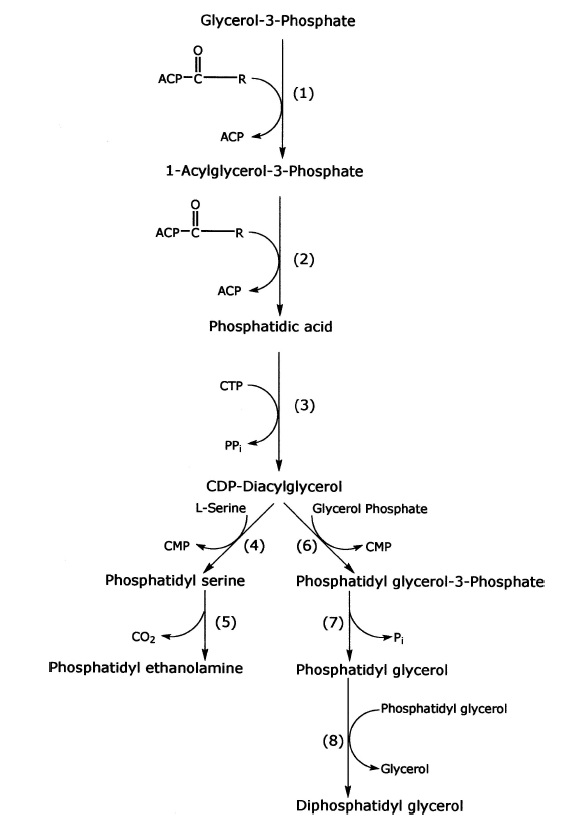
Synthesis of phospholipids in bacteria.
Enzymes for synthesis are as follows:
(1) sn-glycerol-3-phosphate acyltransferase (sn indicates stereospecifically numbered);
(2) 1-acylsn-glycerol-3-phosphate acyltransferase;
(3) phosphatidate cytidylyltransferase;
(4) phosphatidyl serine synthase;
(5) phosphatidyl serine decarboxylase;
(6) glycerol phosphate phosphatidyl transferase;
(7) phosphatidyl glycerol phosphate phosphatase; and
(8 ) diphosphatidyl glycerol synthase. Other abbreviations are as follows: R1 and R2 are fatty acyl groups; ACP is acyl carrier protein.
As in eukaryotic cells, phosphatidic acid PA is the precursor to all the glycerol-based (as distinguished from Lipid A core of lipopolysaccharide (LPS)) phospholipids of E. coli. PA is synthesized in two sequential steps employing long chain acyl-CoA or acyl-ACP (acyl carrier protein) for acylation first at the sn-1 and then the sn-2 position catalyzed by the plsB gene products, respectively.
Abbreviated List of Genes in Bacterial Lipid Metabolism
Gene Protein/Enzyme
aas 2-Acyl-GPE acyltransferase
aasS Acyl-ACP synthetase
accA Carboxyltransferase subunit
accB Biotin carboxy carrier protein
accC Biotin carboxylase
accD Carboxyl transferase subunit
acpP Acyl carrier protein
acpH Acyl carrier protein hydrolase
acpS Acyl carrier protein synthase
cdh CDP-diacylglycerol hydrolase
cdsA CDP-diacylglycerol synthase
cdsS Stabilises mutant CDP-diacylglycerol synthase
cfa Cyclopropane fatty acid synthase
cls Cardiolipin synthase
desA Phospholipid desaturase
desB Acyl-CoA desaturase
desT Transcriptional regulator
desR Transcriptional regulator
dgkA Diacylglycerol kinase (Gram-negative)
dgkB Diacylglycerol kinase (Gram-positive)
fabA β-Hydroxydecanoyl-ACP dehydratase I
fabB β-Ketoacyl-ACP synthase I
fabD Malonyl-CoA:ACP transacylase
fabF β-Ketoacyl-ACP synthase II
fabG β-Ketoacyl-ACP reductase
fabH β-Ketoacyl-ACP-synthase III
fabI Enoyl-ACP reductase I
fabK Enoyl-ACP reductase II
fabL Enoyl-ACP reductase III
fabM trans-2-Enoyl-ACP isomerase
fabN β-Hydroxydecanoyl-ACP dehydrase/isomerase II
fabR Transcriptional regulator (Gram-negative)
fabT Transcriptional regulator (Gram-positive)
fabZ β-Hydroxyacyl-ACP dehydratase
fadA β−Ketoacyl-CoA thiolase
fadB 4-Function enzyme of β-oxidation: β−hydroxyacyl-
CoA dehydrogenase and epimerase; cis-β−trans-2-
enoyl-CoA isomerase and enoyl-CoA hydratase
fadD Acyl-CoA synthetase
fadE Electron transferring flavoprotein
fadF Acyl-CoA dehydrogenase
fadG Acyl-CoA dehydrogenase?
fadH 2,4-Dienoyl-CoA reductase
fadL Long-chain fatty acid transport protein precursor
fadR Transcriptional regulator
fakA The ATP-binding component of fatty acid kinase
fakB The fatty acid-binding component of fatty acid kinase
fapR Transcriptional regulator
fatA Unknown, possible transcription factor
gpsA Glycerol phosphate synthase
htrB KDO2-lipid IVA acyloxy lauroyltransferase
kdtA KDO transferase
lplT Lysophospholipid flippase
lpxA UDP-GlcNAc β-hydroxymyristoyl-ACP acyltransferase
lpxB Disaccharide-1-P synthase
lpxC UDP-β−O-hydroxymyristoyl-GlcNAc deacetylase
lpxD UDP-β−O-hydroxymyristoyl-GlcN N-acyltransferase
lpxK Disaccharide-1-P 4′-kinase
lpxP KDO2-lipid IVA acyloxy palmitoyltransferase
mdoB Phosphatidylglycerol:membrane-oligosaccharide glycerophosphotransferase
msbA Lipid flippase
msbB KDO2-Lipid IVA acyloxy myristoyl transferase
pgpA PGP phosphatase
pgpB PGP phosphatase
pgsA PGP synthase
pldA Detergent-resistant phospholipase A
pldB Inner membrane lysophospholipase
plsB Glycerol phosphate acyltransferase
plsC 1-Acylglycerol phosphate acyltransferase
plsX Acyl-ACP phosphotransferase
plsY Acylphosphate:glycerolphosphate acyltransferase
psd PS decarboxylase
pssA PS synthase
tesA Thioesterase I
tesB Thioesterase II
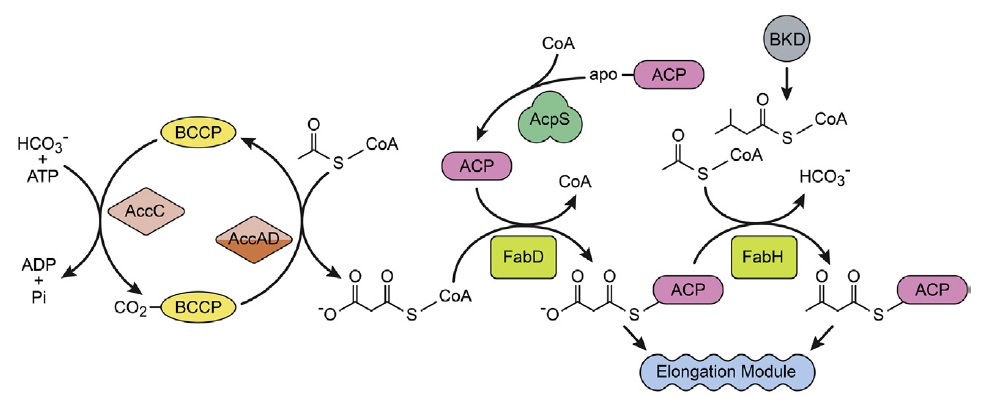
The initiation module.
This group of enzymes is responsible for the initiation of FASII. Acetyl-CoA carboxylase is a 4-protein complex that produces malonyl-CoA by a two-step reaction. AccC carboxylates biotin attached to BCCP, and then AccAD transfers the carboxyl group to acetyl- CoA. Acyl carrier protein (ACP) is the key acyl group carrier in FASII. The primary gene product is an apoprotein that is converted to ACP by the transfer of the 4′-phosphopantetheine moiety of CoA to ACP catalysed by ACP synthase (AcpS). Malonyl-CoA:ACP transacylase (FabD) maintains an equilibrium between the malonyl-CoA and malonyl-ACP pools. FabH is the initiation-condensing enzyme (3-ketoacyl-ACP synthase III) that condenses acetyl-CoA to malonyl-ACP to form the first 3-ketoacyl-ACP intermediate in FASII. Each time FabH fires, a new acyl chain is made. Bacteria that produce branched-chain fatty acids use a branched-chain acyl-CoA primer derived from amino acid metabolism via branched-chain ketoacid dehydrogenase (BKD).
THE INITIATION MODULE
Acyl Carrier Protein
A central feature of fatty acid biosynthesis FASII is that the intermediates are carried from enzyme to enzyme by a small (8.86 kDa), acidic, and highly soluble protein, ACP, the product of the acpP gene. ACP is one of the most abundant proteins in E. coli, constituting about 0.25% of the total soluble protein (∼6 × 104 molecules/cell). The acyl intermediates of fatty acid biosynthesis are bound to the protein through a thioester linkage to the terminal sulphhydryl of the 4′-phosphopantetheine prosthetic group. The prosthetic group contains the only thiol group of ACP and is attached to the protein via a phosphodiester linkage to Ser-36. ACP must interact specifically and transiently with all of the enzymes of FASII and does so through interactions with exposed negatively charged residues on ACP with a patch of positive residues on the surfaces of the fab enzymes. The ACP pool in normally growing cells is approximately one eighth of the coenzyme A (CoA) pool, the other acyl group carrier in cells. The prosthetic group of ACP is derived from CoA, and the common feature of both is the pantetheine arm for thioester formation. ACP is maintained in the active, holoform in vivo, indicating that the supply of prosthetic group does not limit fatty acid biosynthesis. The 4′-phosphopantetheine prosthetic group is transferred from CoA to apo-ACP by the 14 kDa trimeric [ACP]synthase (Figure above). The [ACP]synthase from Bacillus subtilis was crystallized in complex with ACP to give the first detailed look at ACP-protein interactions. ACP is one of the most interactive proteins in bacterial physiology, and the physiological significance of its recognition by all its binding partners continues to be the subject of research. The ACPs from one bacterial species readily substitutes for other ACPs in FASII due to the conservation of residues along helix-2, termed the recognition helix. The conserved negatively charged and hydrophobic residues of ACP interact with a complementary constellation of basic and hydrophobic residues on the target protein that are adjacent to the active site tunnel. Whereas the fab enzymes do not have a conserved primary sequence that can be recognized as an ACP recognition motif, their 3-dimensional structures illustrate that the basic hydrophobic recognition region at the active site entrance is a common structural feature critically important for ACP binding.
https://www.youtube.com/watch?v=IiWpNAbbFBQ
1. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3513495/
2. https://sci-hub.tw/https://www.sciencedirect.com/science/article/abs/pii/S1388198112001898
3. https://en.wikipedia.org/wiki/Glycerol_3-phosphate#:~:text=Glycerol%203%2Dphosphate%20is%20synthesized,cycle%20intermediates%20via%20glyceroneogenesis%20pathway.
In 1956 Kennedy and Weiss published the seminal paper in the phospholipid field that formed the biological foundation for future studies in all organisms thereby establishing the “Kennedy Pathways” for phospholipid synthesis.
There is a large diversity in the structures and metabolism of membrane lipids, and not a single paradigm can be considered typical.
As we now know, phosphatidic acid (PA1, 1,2 diacyl-sn-glycerol-3-phosphate (G3P) ) is the precursor leading to the biosynthesis of the remaining phospholipids in Bacteria and Eukarya.

sn-Glycerol 3-phosphate is a phosphoric ester of glycerol, which is a component of glycerophospholipids. 3
Although the spectrum of phospholipid headgroup structures produced by bacteria is large, the key precursor to all of these molecules is phosphatidic acid (PtdOH). 2 Glycerol-3-phosphate derived from the glycolysis via glycerol-phosphate synthase is the universal source for the glycerol backbone of PtdOH.
My comment: This is another catch22 conundrum. Glycolysis requires a functional membrane to function. But modern membranes depend on membrane precursor phosphatidic acid to be synthesized. Another example of why cells are irreducibly complex and interdependent.
The formation of G3P from the reduction of dihydroxyacetone phosphate by the G3P synthase (GpsA) is the only de novo pathway to G3P in bacteria

Pathways for the biosynthesis of PtdOH in bacteria.
(A) PtdOH metabolism in E. coli is representative of the bacteria that utilize the PlsB/PlsC acyltransferase pathway to PtdOH. These acyltransferases use either acyl-ACP substrates produced by type II fatty acid synthesis (FASII) or acyl-CoA thioesters generated by the activation of exogenous fatty acids by acyl-CoA synthetase (FadD). The PlsB pathway is largely confined to the γ-proteobacteria.
(B) PtdOH metabolism in S. aureus is representative of bacteria that utilize the PlsX/PlsY/PlsC acyltransferase pathway to PtdOH. Acyl-ACP generated by FASII is either used by PlsC to acylate the 2-position of LPA or is converted by PlsX to acyl-PO4 for incorporation into the 1-position by PlsY. This is the only pathway present in most Gram-positive pathogens. In both schemes, the G3P backbone is produced by GpsA, but may also be obtained from the environment by the GlpF/GlpK pathway. Many bacteria also have a transport system for G3P (not shown). Exogenous fatty acids are incorporated into phospholipid following their ligation to ACP by acyl-ACP synthetases (AAS). PlsB, PlsC, PlsY and GlpF are intrinsic membrane proteins. AAS, FadD and PlsX are soluble proteins that are thought to interact with the membrane interface.
There are two distinct families of enzymes responsible for the acylation of the 1-position of glycerol-3-phosphate. The PlsB acyltransferase was discovered in Escherichia coli, and homologs are present in many eukaryotes. This protein family primarily uses acyl–acyl carrier protein (ACP) endproducts of fatty acid synthesis as acyl donors, but may also use acyl-CoA derived from exogenous fatty acids. The second protein family, PlsY, is more widely distributed in bacteria and utilizes the unique acyl donor, acyl-phosphate, which is produced from acyl-ACP by the enzyme PlsX. The acylation of the 2-position is carried out by members of the PlsC protein family. All PlsCs use acyl-ACP as the acyl donor, although the PlsCs of the γ-proteobacteria also may use acyl-CoA. Phospholipid headgroups are precursors in the biosynthesis of other membrane-associated molecules and the diacylglycerol product of these reactions is converted to PtdOH by one of two distinct families of lipid kinases. The central importance of the de novo and recycling pathways to PtdOH in cell physiology suggest that these enzymes are suitable targets for the development of antibacterial therapeutics in Gram-positive pathogens. This article is part of a Special Issue entitled Phospholipids and Phospholipid Metabolism.
Archaea phospholipids differ in that they are composed of sn-glycerol-1-phosphate in ether linkage at the 2- and 3-positions to long-chain poly isoprenoids. 1
Synthesis of phosphatidic acid

Synthesis of phospholipids in bacteria.
Enzymes for synthesis are as follows:
(1) sn-glycerol-3-phosphate acyltransferase (sn indicates stereospecifically numbered);
(2) 1-acylsn-glycerol-3-phosphate acyltransferase;
(3) phosphatidate cytidylyltransferase;
(4) phosphatidyl serine synthase;
(5) phosphatidyl serine decarboxylase;
(6) glycerol phosphate phosphatidyl transferase;
(7) phosphatidyl glycerol phosphate phosphatase; and
(8 ) diphosphatidyl glycerol synthase. Other abbreviations are as follows: R1 and R2 are fatty acyl groups; ACP is acyl carrier protein.
As in eukaryotic cells, phosphatidic acid PA is the precursor to all the glycerol-based (as distinguished from Lipid A core of lipopolysaccharide (LPS)) phospholipids of E. coli. PA is synthesized in two sequential steps employing long chain acyl-CoA or acyl-ACP (acyl carrier protein) for acylation first at the sn-1 and then the sn-2 position catalyzed by the plsB gene products, respectively.
Abbreviated List of Genes in Bacterial Lipid Metabolism
Gene Protein/Enzyme
aas 2-Acyl-GPE acyltransferase
aasS Acyl-ACP synthetase
accA Carboxyltransferase subunit
accB Biotin carboxy carrier protein
accC Biotin carboxylase
accD Carboxyl transferase subunit
acpP Acyl carrier protein
acpH Acyl carrier protein hydrolase
acpS Acyl carrier protein synthase
cdh CDP-diacylglycerol hydrolase
cdsA CDP-diacylglycerol synthase
cdsS Stabilises mutant CDP-diacylglycerol synthase
cfa Cyclopropane fatty acid synthase
cls Cardiolipin synthase
desA Phospholipid desaturase
desB Acyl-CoA desaturase
desT Transcriptional regulator
desR Transcriptional regulator
dgkA Diacylglycerol kinase (Gram-negative)
dgkB Diacylglycerol kinase (Gram-positive)
fabA β-Hydroxydecanoyl-ACP dehydratase I
fabB β-Ketoacyl-ACP synthase I
fabD Malonyl-CoA:ACP transacylase
fabF β-Ketoacyl-ACP synthase II
fabG β-Ketoacyl-ACP reductase
fabH β-Ketoacyl-ACP-synthase III
fabI Enoyl-ACP reductase I
fabK Enoyl-ACP reductase II
fabL Enoyl-ACP reductase III
fabM trans-2-Enoyl-ACP isomerase
fabN β-Hydroxydecanoyl-ACP dehydrase/isomerase II
fabR Transcriptional regulator (Gram-negative)
fabT Transcriptional regulator (Gram-positive)
fabZ β-Hydroxyacyl-ACP dehydratase
fadA β−Ketoacyl-CoA thiolase
fadB 4-Function enzyme of β-oxidation: β−hydroxyacyl-
CoA dehydrogenase and epimerase; cis-β−trans-2-
enoyl-CoA isomerase and enoyl-CoA hydratase
fadD Acyl-CoA synthetase
fadE Electron transferring flavoprotein
fadF Acyl-CoA dehydrogenase
fadG Acyl-CoA dehydrogenase?
fadH 2,4-Dienoyl-CoA reductase
fadL Long-chain fatty acid transport protein precursor
fadR Transcriptional regulator
fakA The ATP-binding component of fatty acid kinase
fakB The fatty acid-binding component of fatty acid kinase
fapR Transcriptional regulator
fatA Unknown, possible transcription factor
gpsA Glycerol phosphate synthase
htrB KDO2-lipid IVA acyloxy lauroyltransferase
kdtA KDO transferase
lplT Lysophospholipid flippase
lpxA UDP-GlcNAc β-hydroxymyristoyl-ACP acyltransferase
lpxB Disaccharide-1-P synthase
lpxC UDP-β−O-hydroxymyristoyl-GlcNAc deacetylase
lpxD UDP-β−O-hydroxymyristoyl-GlcN N-acyltransferase
lpxK Disaccharide-1-P 4′-kinase
lpxP KDO2-lipid IVA acyloxy palmitoyltransferase
mdoB Phosphatidylglycerol:membrane-oligosaccharide glycerophosphotransferase
msbA Lipid flippase
msbB KDO2-Lipid IVA acyloxy myristoyl transferase
pgpA PGP phosphatase
pgpB PGP phosphatase
pgsA PGP synthase
pldA Detergent-resistant phospholipase A
pldB Inner membrane lysophospholipase
plsB Glycerol phosphate acyltransferase
plsC 1-Acylglycerol phosphate acyltransferase
plsX Acyl-ACP phosphotransferase
plsY Acylphosphate:glycerolphosphate acyltransferase
psd PS decarboxylase
pssA PS synthase
tesA Thioesterase I
tesB Thioesterase II

The initiation module.
This group of enzymes is responsible for the initiation of FASII. Acetyl-CoA carboxylase is a 4-protein complex that produces malonyl-CoA by a two-step reaction. AccC carboxylates biotin attached to BCCP, and then AccAD transfers the carboxyl group to acetyl- CoA. Acyl carrier protein (ACP) is the key acyl group carrier in FASII. The primary gene product is an apoprotein that is converted to ACP by the transfer of the 4′-phosphopantetheine moiety of CoA to ACP catalysed by ACP synthase (AcpS). Malonyl-CoA:ACP transacylase (FabD) maintains an equilibrium between the malonyl-CoA and malonyl-ACP pools. FabH is the initiation-condensing enzyme (3-ketoacyl-ACP synthase III) that condenses acetyl-CoA to malonyl-ACP to form the first 3-ketoacyl-ACP intermediate in FASII. Each time FabH fires, a new acyl chain is made. Bacteria that produce branched-chain fatty acids use a branched-chain acyl-CoA primer derived from amino acid metabolism via branched-chain ketoacid dehydrogenase (BKD).
THE INITIATION MODULE
Acyl Carrier Protein
A central feature of fatty acid biosynthesis FASII is that the intermediates are carried from enzyme to enzyme by a small (8.86 kDa), acidic, and highly soluble protein, ACP, the product of the acpP gene. ACP is one of the most abundant proteins in E. coli, constituting about 0.25% of the total soluble protein (∼6 × 104 molecules/cell). The acyl intermediates of fatty acid biosynthesis are bound to the protein through a thioester linkage to the terminal sulphhydryl of the 4′-phosphopantetheine prosthetic group. The prosthetic group contains the only thiol group of ACP and is attached to the protein via a phosphodiester linkage to Ser-36. ACP must interact specifically and transiently with all of the enzymes of FASII and does so through interactions with exposed negatively charged residues on ACP with a patch of positive residues on the surfaces of the fab enzymes. The ACP pool in normally growing cells is approximately one eighth of the coenzyme A (CoA) pool, the other acyl group carrier in cells. The prosthetic group of ACP is derived from CoA, and the common feature of both is the pantetheine arm for thioester formation. ACP is maintained in the active, holoform in vivo, indicating that the supply of prosthetic group does not limit fatty acid biosynthesis. The 4′-phosphopantetheine prosthetic group is transferred from CoA to apo-ACP by the 14 kDa trimeric [ACP]synthase (Figure above). The [ACP]synthase from Bacillus subtilis was crystallized in complex with ACP to give the first detailed look at ACP-protein interactions. ACP is one of the most interactive proteins in bacterial physiology, and the physiological significance of its recognition by all its binding partners continues to be the subject of research. The ACPs from one bacterial species readily substitutes for other ACPs in FASII due to the conservation of residues along helix-2, termed the recognition helix. The conserved negatively charged and hydrophobic residues of ACP interact with a complementary constellation of basic and hydrophobic residues on the target protein that are adjacent to the active site tunnel. Whereas the fab enzymes do not have a conserved primary sequence that can be recognized as an ACP recognition motif, their 3-dimensional structures illustrate that the basic hydrophobic recognition region at the active site entrance is a common structural feature critically important for ACP binding.
https://www.youtube.com/watch?v=IiWpNAbbFBQ
1. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3513495/
2. https://sci-hub.tw/https://www.sciencedirect.com/science/article/abs/pii/S1388198112001898
3. https://en.wikipedia.org/wiki/Glycerol_3-phosphate#:~:text=Glycerol%203%2Dphosphate%20is%20synthesized,cycle%20intermediates%20via%20glyceroneogenesis%20pathway.
Last edited by Admin on Fri Jul 17, 2020 5:46 pm; edited 2 times in total


