Mechanisms and Controls of DNA Replication in Bacteria 1
IntroductionDNA is the polymeric molecule that contains all the genetic information in a cell.
This genetic information encodes the instructions to make a copy of itself, for the cellular structure, for the operative cellular machinery and also contains the regulatory signals, which determine when parts of this machinery should be on or off. The operative machinery in turn, is responsible for the cells functions either metabolically or in interactions with the environment. Part of this cellular machinery devoted to
DNA metabolism is responsible for
DNA replication,
DNA-repair and for the regulation of gene expression. In this chapter we will focus our discussion on the mechanisms and controls that conduct
DNA replication in bacteria, including the components, functions and regulation of
replication machinery. Most of our discourse will consider this biological process in Escherichia coli but when possible we will compare it to other bacterial models, mainly Bacillus subtilis and Caulobacter crescentus as examples of organisms with asymmetrical cell division. In order to maintain a bacterial population it is necessary that cells divide, but before the physical division of a daughter cell from its mother, it is necessary among other check points, that the
DNA has been replicated accurately. This is done by the universal semiconservative
replication process of
DNA-strands, which generates two identical strand copies from their parent templates. To better understand this process it has been divided into three phases: initiation, elongation and termination of
DNA replication. In each of these steps, multiple stable and transient interactions are involved and we have summarized them below.
The study of the cell-cycle in bacteria is usually divided into three stages: the period between cell-division (cell birth) and the initiation of chromosome
replication, the period required to complete
DNA replication (elongation of
DNA) and, the final phase, which goes from the end of
DNA replication until the completion of celldivision (Wang & Levine, 2009). Under the best growing-conditions,
DNA replication starts immediately after cell division in most cells (Wang et al., 2005). Since
replication of the chromosome takes more time than that the necessary for cell division under optimal culture conditions, such as E. coli growing in rich media, at 37ºC with good aeration, it can happen that more than one event of
DNA replication can occur per cell cycle (Zakrzewska-Czerwinska et al., 2007). For the purposes of this work we shall divide the
DNA replication process in bacteria into three steps: initiation, elongation and termination as follows.
3. Regulation of DNA replication The regulation of DNA replication is a vital cellular process. In a general view, DNA replication is controlled by a series of mechanisms that are centered on the control of cellular DnaA levels, its availability as a free protein and modulation of its activity by binding the small-molecule ligand ATP (Leonard & Grimwade, 2009); the other point of control is by modulating the accessibility of replisome components to the oriC region on the
DNA. We discuss some aspects of these regulatory mechanisms below.


 3.1 Regulatory mechanisms of DNA replication in E. coli
3.1 Regulatory mechanisms of DNA replication in E. coli One of the main mechanisms associated with
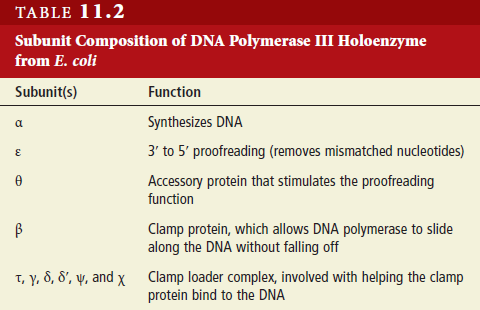
DNA replication is the so-called RIDA system (Regulatory Inactivation of DnaA). The elements of this system are the sliding-clamp of
DNA polymerase III and Hda (Homologous to DnaA). This mechanism takes place when DnaA is activated by its binding to ATP. The accumulation of DnaA in this active form leads to the initiation of chromosomal
replication since it facilitates its binding to the oriC on the
DNA. DnaA reverts to its inactive form DnaA-ADP by hydrolysis of ATP (Katayama et al., 1998). Hda-ADP is the monomeric active form for promoting the hydrolysis of
DNA-ATP, a process which is mediated by the slider-loader clamp (Su’etsugu et al., 2008).
This inactivating regulation of DnaA is key for preventing the over-initiation of replicative events during the cell cycle (Katayama & Sekimizu, 1999). The free-living bacteria C. crescentus also presents this regulatory mechanism, as it has HdaA, a protein similar to the E. coli Hda. In C. crescentus HdaA also inactivates DnaA in a
replication-coordinated manner, if
DNA replication is successfully initiated then HdaA and the ┚-sliding clamp promote the hydrolysis of DnaA-ATP to DnaA-ADP and force DnaA to leaves the oriC (Collier & Shapiro, 2009). A conserved bacterial protein, YabA, has been found in B. subtilis and other Gram-positive bacteria where it acts as a repressor for initiation of
DNA replication. This is achieved by forming a complex with DnaA and the ┚-sliding clamp independently of the
DNA, a common activity shared between Hda and YabA (Mott & Berger, 2007). Thus the RIDA system is present in B. subtilis and is also the primary mechanism for regulation of
DNA replication in this bacterium (Noirot-Gros et al., 2006). The formation of the oriC and DnaA complex is assisted by the protein DiaA, which forms homo-tetramers and binds various DnaA molecules, especially in the active form of DnaA-ATP but it can also stimulate the formation of the DnaA-ADP-oriC complex, this is an inactive complex for initiation of
replication (Ishida et al., 2004).
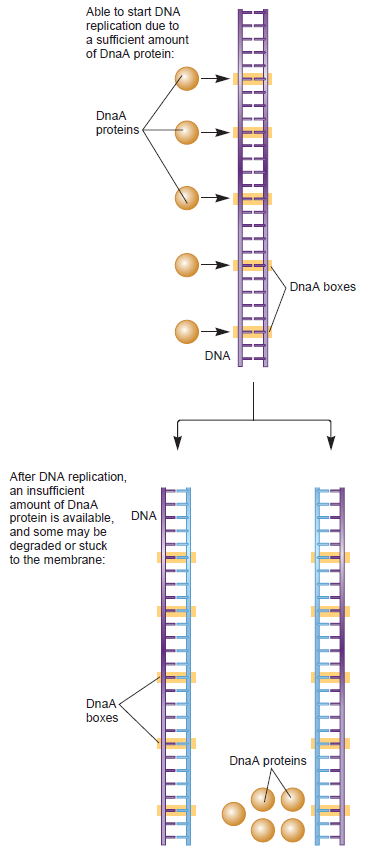
Another mechanism that regulates the initiation of
DNA replication is by controlling the availability of free DnaA to bind to DnaA boxes on the oriC . Here the role of the 1kb datA locus, which is localized near (downstream) from the oriC is important. The datA locus shows high affinity for DnaA, even more than the DnaA boxes on the oriC. Thus the datA region is able to bind over 300 DnaA molecules whereas oriC binds to 45 DnaA monomers (Kitagawa et al., 1998). The operability of this mechanism is facilitated by the fact that the oriC had only few DnaA boxes compared to the datA locus and by the close proximity of data in respect to oriC on the
DNA molecule (Figure 6).
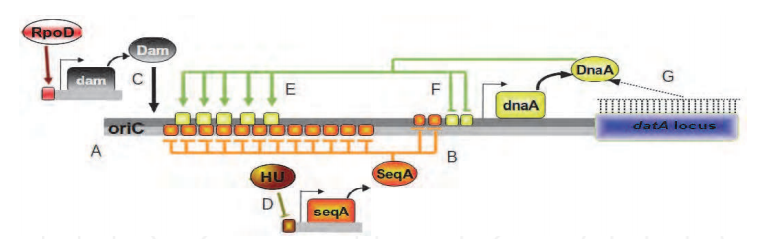 Fig. 6. Mechanisms that regulate DNA replication in E. coli. A) The newly replicated DNA duplex is in a hemimethylated state. B) SeqA binds to the hemimethylated GATC sites immediately after they are replicated. C) RpoD activates the transcription of dam and Dam methylates GATC sites of the newly synthesized strand. D) HU represses the transcription of SeqA. E) DnaA binds to the DnaA boxes on the oriC region. F) when there are many DnaA molecules they repress the transcription of the dnaA gene. G) datA locus binds many DnaA molecules.
Fig. 6. Mechanisms that regulate DNA replication in E. coli. A) The newly replicated DNA duplex is in a hemimethylated state. B) SeqA binds to the hemimethylated GATC sites immediately after they are replicated. C) RpoD activates the transcription of dam and Dam methylates GATC sites of the newly synthesized strand. D) HU represses the transcription of SeqA. E) DnaA binds to the DnaA boxes on the oriC region. F) when there are many DnaA molecules they repress the transcription of the dnaA gene. G) datA locus binds many DnaA molecules. One related control system depends on the property of DnaA to act as a transcription factor and to the presence of DnaA boxes in the promoter regions of several genes. In most cases DnaA represses the expression of the associated gene but in some cases it can activates certain genes (Messer & Weigel, 1997). DnaA regulates around 10 genes in E. coli as documented in RegulonDB (Gama-Castro et al., 2010). The transcription of DnaA is one of the most important regulatory mechanisms that directly affect the
replication of
DNA and one of the proteins that negatively regulate the expression of dnaA is DnaA itself (Figure 6). At high levels DnaA binds to the DnaA boxes in the promoter region and impedes transcription. This auto-repressive process directly affects the amount of DnaA-ATP available and controls the efficiency of initiation of
DNA replication (Mott & Berger, 2007). In C. crescentus, it was found that DnaA also auto-represses the transcription of its own gene but additionally DnaA is highly unstable in this organism and gradually degrades after initiating a
replication event (Gorbatyuk & Marczynski, 2005).
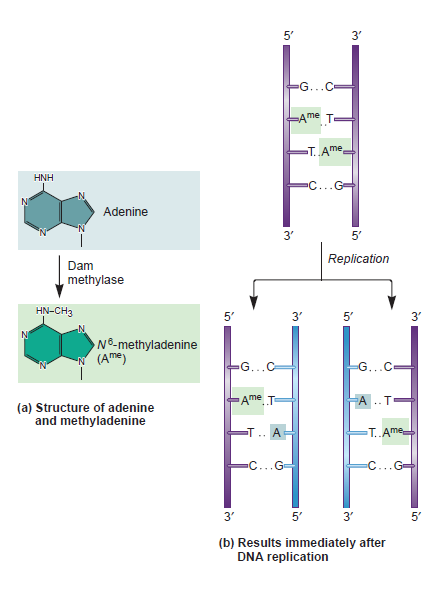
3.2 Regulation of DNA replication by DNA methylation A requirement for initiation of
DNA replication is that both
DNA strands are methylated, principally the adenine nucleotide in the GATC motifs, this process is mediated by Dam (
DNA adenine methyltranferase), (Wion & Casadésus, 2006). Dam binds to the
DNA nonspecifically, and methylates the GATC motifs (Figure 6). On
DNA strands recently synthesized these motifs are rapidly methylated and exist in the hemimethylated state only during a fraction of the time needed for the
replication of the entire
DNA (Casadésus & Low, 2006). The methylation process occurs asynchronously on the newly synthesized strands; i. e. methylation on the lagging arm occurs only after the ligation of the Okazaki fragments. It is postulated that Dam is always present in a complex bound near the
replication origin, thus the methylation of nascent
DNA strands occurs as soon as polymerization begins. In summary, the presence of hemimethylated GATC sites provides a cue to indicate that
DNA replication has just occurred (Stancheva et al., 1999). Another way to repress the transcription of dnaA is that which occurs immediately after the initiation of
DNA replication. Here, SeqA binds to the hemimethylated GATC sequences in the regulatory regions of the dnaA gene (Lu et al., 1994; Brendler et al., 2000). Similarly, SeqA also represses the
replication of
DNA by binding to the hemimethylated GATC sequence at the oriC, this is possible because SeqA
DNA-binding sites overlap with those of low affinity for DnaA (DnaA boxes) on the oriC. This overlap impedes the complete access of DnaA-ATP to the oriC (Han et al., 2004).
This prevention of replication, dependant of DNA methylation, has been considered as an epigenetic regulatory mechanism because it depends on the chemical modification of the nucleotide residues of the DNA and not in its sequence. ConclusionsThe replication of DNA is a complex process in which a great number of regulators and mechanisms are involved, one of the most important is the DnaA protein. Replication normally begins by the formation of a complex of DnaA at the oriC region, with the assistance of DiaA, and the incorporation of some proteins that form the replisome, subsequently the formation of the open complex takes place, followed by a complex interaction of the proteins needed to execute and complete the
DNA replication. The process finalizes with the recognition of the ter site and disassembly of the replisome. Many of the proteins are broadly conserved within the bacteria but some special factors are required in bacteria which undergo particular processes such as asymmetrical cell division.
In general these processes are controlled by a series of circuits, which usually center on the oriC and affect the activity of DnaA. The result is regulation of the initiation step of DNA replication. Some of the regulatory mechanisms are time-dependent allowing only one
DNA replication event per cell cycle. The methylation state of the
DNA-strands is another important condition that not only controls the possibility of starting
DNA replication but also regulates the transcription of many genes important for the execution of this function. All or certain of these mechanisms are adjusted under some special conditions, such as when the stringent response is triggered by amino acid starvation. In some bacteria with extremely reduced genomes it is still a mystery as to how
DNA replication takes place and how it is controlled. Many of these latter organisms lack several important proteins implicated in the control and execution of
DNA replication, and these bacteria can be useful as models for generating a system with the minimal components necessary for
DNA replication.
1) http://cdn.intechopen.com/pdfs-wm/20627.pdf